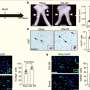
To better understand the signaling pathways active in sarcomas, researchers at Moffitt Cancer Center used state-of-the-art mass spectrometry-based proteomics to characterize a family of protein enzymes that act as "on" or "off" switches important in the biology of cancer. The tyrosine kinases they identified, the researchers said, could act as "drivers" for the growth and survival of sarcomas.
Sarcomas are relatively rare forms of cancer. In contrast to carcinomas, which arise from epithelial cells (in breast, colon and lung cancers, for example), sarcomas are tumors derived from bone, fat, muscle or vascular tissues.
"Sarcomas are rare, diverse malignancies that arise from connective tissues," said study lead author Eric B. Haura, M.D., program leader for Experimental Therapeutics. "We hypothesized that we could identify important proteins that drive the growth and survival of these poorly understood sarcomas using an approach to characterize signaling proteins using mass spectrometry."
According to Haura, whose lab focuses on signaling pathways in cancer and understanding the role of kinases, protein phosphorylation plays a significant role in a wide range of cellular processes and is commonly disrupted in diseases such as cancer. The study approach is novel by engaging proteomics, an emerging and increasingly powerful approach to study proteins in disease in a more global and unbiased manner.
In this study, the Moffitt researchers identified 1,936 unique tyrosine phosphorylated peptides corresponding to 844 unique phospho-tyrosine proteins and found 39 tyrosine kinases in sarcoma cells. Of the 99 tyrosine kinases present in the human genome, the research team identified peptides corresponding to nearly 40 percent of the tyrosine kinome.
"Tyrosine kinases play an important role in controlling the hallmarks of cancer, and they have a proven track record as druggable targets for cancer treatment. Our goal was to produce a 'landscape' of tyrosine phosphorylated proteins and tyrosine kinases prioritized for subsequent functional validation," Haura said. "In our study, we identified numerous tyrosine kinases that can be important for cellular signaling in human sarcomas that could drive the natural progression of sarcoma and, therefore, could be targeted by small molecule inhibitors aimed at altering sarcoma progression."
Questions remain, however, about which, if any, of the 40 tyrosine kinases the researchers identified in sarcoma tumor cell lines act to regulate sarcoma tumor cell growth and tumor survival.
"The answers to this question can help prioritize which potential targets to examine further, including advancement into trials of patients with advanced sarcoma," explained Haura. "As a first step, we screened sarcoma cell lines against a number of inhibitors selected, all based on the tyrosine kinases we identified, and identified some active drugs."
While the researchers found kinases in sarcoma cells that deserved further study, they also concluded that the sarcoma cells tested expressed multiple tyrosine kinases. That great number may limit the effectiveness of targeted agents.
"We think this approach could hold promise in profiling tumors directly from patients and can complement existing genetic data on sarcomas. Our results show this is feasible in tumor tissues, and we hope to advance this further by directly studying additional tumors from sarcoma patients."
Their study appeared in a recent issue of Cancer Research published by the American Association for Cancer Research.