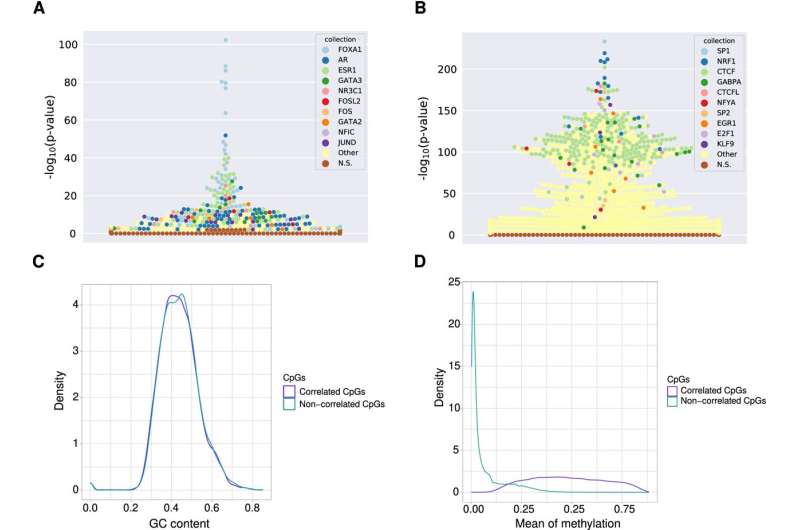
TF binding signatures at FOXA1-associated emCpGs versus de novo methylation-protected CpGs in breast cancer. A Beeswarm plot depicting TFBS sets enrichment (y-axis) specific to regions surrounding emCpGs associated with FOXA1. Each point corresponds to a TFBS data set in UniBind (one color per TF, see legend). B Beeswarm plot depicting TFBS sets enrichment (y-axis) specific to regions surrounding non-correlated CpGs, which are close to FOXA1 TFBSs but whose DNA methylation levels do not correlate with FOXA1 expression in breast cancer samples. C Density distribution (y-axis) of GC contents (x-axis) at regions surrounding FOXA1 emCpGs (purple) and non-correlated CpGs (green). D Density distribution (y-axis) of mean methylation levels (x-axis) across breast cancer samples for FOXA1 emCpGs (purple) and non-correlated CpGs (green). Credit: Epigenetics & Chromatin (2022). DOI: 10.1186/s13072-022-00444-9
Reducing the burden of cancer in individuals and society is a major area of concern globally. In Norway, by the end of 2020, 35,515 people were newly diagnosed with cancer and more than 300,000 people were already living with a diagnosis according to the Cancer Registry of Norway. Developing knowledge about the causes of cancer and the underlying processes represents a critical public health priority.
At the Center for Molecular Medicine Norway (NCMM), researchers have investigated 19 cancer types to identify the potential molecular players that regulate aberrant DNA methylation, an epigenetic modification on the DNA associated with the regulation of gene expression. They observed an association between the expression of 13 transcription factors (TFs), which are special genes that bind to the DNA at TF binding sites (TFBSs) to control gene expression, and DNA methylation levels in the vicinity of their TFBSs. Each of these 13 TFs is selected based on their association with changes in DNA methylation levels in at least two cancer types. Their results are published in Epigenetics & Chromatin.
The link between transcription factors and DNA methylation in cancer samples
Within the human body, there are countless genetic processes taking place involving our DNA to allow for healthy development. One of these processes is called DNA methylation—a mechanism that is associated with the regulation of gene expression. Abnormal gene expression patterns are linked to several diseases, including cancer.
"DNA methylation is a natural process associated with the control of gene expression. In a normal case you have a certain pattern of DNA methylation but this can be hijacked by cancer cells and as a result show different (abnormal) patterns," explains researcher Dr. Roza Berhanu Lemma.
Dr. Roza Berhanu Lemma, a researcher at NCMM in the research group of Dr. Anthony Mathelier, and her colleagues have been investigating the interplay between DNA methylation patterns and transcription factor gene expression in cancer samples.
"Abnormal DNA methylation patterns are frequently reported in cancer patients. We wanted to find out what the mechanism of this dysregulation is and to see if there are transcription factors involved in modulating this process," Lemma explained.
Researchers found 13 special genes associated with DNA methylation patterns in cancer samples
Lemma and her colleagues developed a computational framework to perform statistical analyses on the relationship between transcription factors and DNA methylation using patient specific data from The Cancer Genome Atlas, along with DNA binding information for human transcription factors from UniBind, a database they developed.
They found 13 transcription factors whose expression levels were correlated with the amount of DNA methylation around their binding sites on the DNA. Among these 13 transcription factors, most of them are "pioneer factors," which means that they have a special ability to bind to regions of the DNA that are otherwise inaccessible and methylated.
Potential of the team's finding
The team's finding has the potential to lead to better cancer diagnosis and/or prognosis.
"We identified special properties between these transcription factors and the DNA methylation patterns close to the binding sites of the transcription factors. One approach could be to use the DNA methylation patterns that are regulated by these transcription factors as potential biomarkers," Lemma explained.
"Through this type of research we contribute additional knowledge so that in the future we have the possibility to utilize distinct epigenetic profiles of the cancers to explore possibilities for potential therapeutic approaches," she says.
This study provides supporting evidence for the involvement of specific genes in the alteration of an important epigenetic mark in cancer. Such discoveries can be paramount to highlight the biological aspects of tumor behavior, function, and identity. Nevertheless, future investigations will be necessary to further understand how this process is associated with the rewiring of gene regulatory networks in cancer cells.
More information: Roza Berhanu Lemma et al, Pioneer transcription factors are associated with the modulation of DNA methylation patterns across cancers, Epigenetics & Chromatin (2022). DOI: 10.1186/s13072-022-00444-9
Provided by University of Oslo