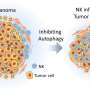
Study authors Adam Getzler, Dapeng Wang and Matthew Pipkin of The Scripps Research Institute collaborated with scientists at the University of California, San Diego. Credit: The Scripps Research Institute
During infection or tumor growth, a type of specialized white blood cells called CD8+ T cells rapidly multiply within the spleen and lymph nodes and acquire the ability to kill diseased cells. Some of these killer T cells then migrate where required to vanquish the germs or cancers.
But how do killer T cells "learn" to leave their home base and amass within specific tissues like the skin, gut, and lung, or solid tumors? Finding the factors that cause T cells to function beyond the lymphoid system and in sites of infection or cancer has proven a tough challenge, but it's essential for developing cancer-fighting immunotherapy strategies.
Writing in the journal Nature, researchers from The Scripps Research Institute and the University of California, San Diego report the discovery that a protein called "Runx3" programs killer T cells to establish residence in tumors and infection sites.
"Runx3 works on chromosomes inside killer T cells to program genes in way that enables the T cells to accumulate in a solid tumor," said Matthew Pipkin, Ph.D., associate professor in the Department of Immunology and Microbiology on the Florida campus of The Scripps Research Institute.
The paper, "Runx3 programs CD8+ T cell residency in non-lymphoid tissues and tumors," appears in Nature's Dec. 14 issue.
There are two main strategies in cancer immunotherapy that employ killer T cells, Pipkin said. Checkpoint inhibitor blockade unleashes killer T cells, prompting them to accumulate in tumors more aggressively. Adoptive cell transfer, meanwhile, involves re-infusing a patient's own immune cells after they have been engineered in the lab to recognize and destroy the patient's specific cancer.
The adoptive cell transfer strategy has worked stunningly well in some blood cancers associated with the lymphoid system, so far. But there appears to be less efficient activity of T cells in solid tumors, Pipkin said.
"The gene programs and signals for how the T cells take up residence in tissues outside of the general circulation was not really well understood," Pipkin said.
To discover factors that control T cell residency beyond the lymphoid system, Pipkin's team worked collaboratively with the laboratory of UC San Diego's Ananda Goldrath, who compared the gene expression of CD8+ T cells found in non-lymphoid tissue to those found in the general circulation. From a list of potential factors, they employed an RNA interference screening strategy which can test the actual function of thousands of factors simultaneously. Pipkin's lab had developed the screening strategy in collaboration with Shane Crotty at the La Jolla Institute for Allergy and Immunology.
"We found a distinct pattern," Pipkin said. "The screens showed that Runx3 is one at the top of a list of regulators essential for T cells to reside in nonlymphoid tissues." Moreover, Runx3 was able to engage a specific gene program that is found in natural tissue-resident and tumor infiltrating CD8+ T cells, he said.
The group further assessed whether Runx3 had a role in directing white blood cells that attack solid tumors in mouse melanoma models. They found that adoptive cell transfer of cancer-specific killer T cells that overexpressed Runx3 delayed tumor growth and prolonged survival, while mouse models treated with those lacking Runx3 fared much worse than normal.
"If we enhance Runx3 activity in the cells, the tumors are significantly smaller and there is greater survival compared to the control group," Pipkin said.
Knowing that modulating Runx3 activity in T cells influences their ability to reside in solid tumors opens new opportunities for improving cancer immunotherapy, Pipkin said.
"The upshot is we could probably use Runx3 to reprogram adoptively transferred cells to help drive them to amass in solid tumors," he said. He added that a collaboration of specialists energized the research. "It was a fantastic collaboration, it all came together very quickly," Pipkin said.
More information: J. Justin Milner et al, Runx3 programs CD8+ T cell residency in non-lymphoid tissues and tumours, Nature (2017). DOI: 10.1038/nature24993
Journal information: Nature
Provided by The Scripps Research Institute