Genomic screen nets hundreds of human proteins exploited by HIV
In some ways, HIV resembles a minimalist painter, using a few basic components to achieve dramatic effects. The virus contains just nine genes encoding 15 proteins, which wreak havoc on the human immune system. But this bare bones approach could have a fatal flaw. Lacking robust machinery, HIV hijacks human proteins to propagate, and these might represent powerful therapeutic targets.
Using a technique called RNA interference to screen thousands of genes, Harvard Medical School researchers have now identified 273 human proteins required for HIV propagation. The vast majority had not been connected to the virus by previous studies. The work appears online in Science Express on Jan. 10.
Drugs currently used to treat the viral infection interact directly with the virus itself, and it’s quite simple for the rapidly mutating virus to avoid destruction by altering how it interacts with these chemicals. Patients use a cocktail of HIV inhibitors because the virus is less likely to evolve resistance to multiple drugs at the same time. But some HIV strains have still managed to evade particular drugs. These could eventually develop resistance to several drugs, especially among patients who don’t adhere to their regimens.
“Antiviral drugs are currently doing a good job of keeping people alive, but these therapeutics all suffer from the same problem, which is that you can get resistance, so we decided to take a different approach centered on the human proteins exploited by the virus,” says Harvard Medical School (HMS) Professor and senior author Stephen Elledge, who holds primary appointments in the Department of Genetics and at Brigham and Women’s Hospital. “The virus would not be able to mutate to overcome drugs that interact with these proteins.”
Labs around the world have made impressive contributions to our understanding of the HIV life cycle. Over the last two decades, they’ve identified dozens of human proteins, or host factors, required for HIV propagation. The new study builds on this work, essentially quadrupling the list of host factors to include proteins involved with a surprising array of cellular functions ranging from protein trafficking to a type of programmed cell death called autophagy.
“The expanded list is a hypothesis generation machine,” explains Elledge, who is also a member of the HMS-Partners Health Care Center for Genetics and Genomics and investigator with the Howard Hughes Medical Institute. “Scientists can look at the list, predict why HIV needs a particular protein, and then test their hypothesis.” He hopes that such research will lead to new therapeutics.
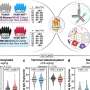
To create the list, postdoctoral researcher and first author Abraham Brass—working with Derek Dyxkhoorn and Nan Yan from HMS Professor Judy Lieberman’s lab—began with a library of short interfering RNAs (siRNAs) targeting specific human genes. Each siRNA disrupts the gene’s ability to produce a particular protein.
With the help of the staff at the Institute of Chemistry and Cell Biology at Longwood (ICCB-L), Brass placed the siRNAs on thousands of human cells, with just one gene being targeted in each well of cells. Thus each well contained cells lacking a particular protein. Next, he unleashed HIV on the cells. If HIV replication was inhibited in a given well, it suggested the missing protein was involved.
Of the 273 proteins he identified, just 36 had been previously implicated in the HIV life cycle. He picked three of the other 237 proteins, and subjected them to a host of careful genetic experiments, proving they too truly play a role in HIV propagation.
Immune cells—the very cells HIV attacks—contain high concentrations of many of the 273 host factors, offering further proof of the list’s validity.
“We’re closing in on a systems level understanding of HIV, which opens new therapeutic avenues,” says Elledge. “We might be able to tweak various parts of the system to disrupt viral propagation without making our own cells sick.”
“This is the first whole genome screen for human proteins required by HIV, and we’re confident that it netted real results,” adds Brass. “Given the method, we missed some proteins, but the majority of the ones we found are highly likely to play a role in HIV propagation.”
Source: Harvard Medical School