New blood test for Down syndrome
Howard Hughes Medical Institute researchers have developed a new prenatal blood test that accurately detected Down syndrome and two other serious chromosomal defects in a small study of 18 pregnant women. If confirmed in larger trials, they say, the test would offer a safer and faster alternative to invasive prenatal tests such as amniocentesis that pose a small risk of miscarriage.
Researchers have long known that a pregnant woman's blood contains small amounts of DNA from the fetus. Howard Hughes Medical Institute researcher Stephen R. Quake and colleagues at Stanford University devised an ingenious way to the scan fetal DNA present in the mother's blood to determine whether the fetus' cells contain extra chromosomes associated with several types of severe birth detects.
The test developed by Quake's team was more accurate than techniques used in previous efforts to diagnose aneuploidy by analyzing fetal DNA. Aneuploidy occurs when there are either too many or too few chromosomes in cells. Down syndrome, for example, is caused by a trisomy -- three copies instead of two -- of chromosome 21.
"We believe this is the first demonstration of a universal, noninvasive test for Down and other aneuploidies," said Quake, senior author of the research article, which was published online in the early edition of the Proceedings of the National Academy of Sciences (PNAS) on October 6, 2008. "We need a larger clinical study to understand a bit more about the best way to implement it, but I am highly optimistic it will be used as a diagnostic test in short order."
Amniocentesis and chorionic villus sampling (CVS), which are currently considered the "gold standard" in prenatal testing, involve sampling cells in amniotic fluid. Those cells are obtained by inserting a needle into the mother's uterus. The procedure carries up to a one percent risk of inducing a miscarriage. For that reason, routine use of these invasive diagnostic tests has largely been limited to women age 35 and older, where the age-related risk of fetal aneuploidy outweighs the risk of miscarriage. As a result, more infants with chromosomal defects are now being born to younger women. Researchers have been working for years to devise noninvasive screening tests that would be safe for that group of women.
In addition, it takes two or three weeks following the amniocentesis or CVS procedures to culture the cells and study the chromosomes. That amount of time can seem like an eternity for anxious parents waiting for the test results, Quake said.
With the new test, scientists only need to draw a small amount of blood from the woman. The blood is analyzed and the results are available within a couple of days, said Quake, who is a bioengineer at Stanford.
Quake's interest in developing new technology to diagnose aneuploidies was sparked when he read a research article published in the journal Science in 2005. That article discussed new methods of noninvasive prenatal testing that relied on measurements of DNA in fetal cells present in the mother's blood. The report said that the low prevalence of fetal cells – only about one in one million maternal cells – made it difficult to isolate enough fetal cells to test for chromosomal abnormalities.
At the time, some researchers were trying to isolate fetal "cell-free DNA" from the mother's blood. Cell-free DNA is fragmented double-stranded DNA that is in the debris of dying fetal cells. Isolating this DNA was a good idea, Quake said, but recovering the vanishingly small amount of fetal cell-free DNA remained a challenge.
After reading the Science article, Quake thought to himself, 'A lot of my work is about counting molecules; this is one problem I know how to do.' He had a brainstorm: Where other researchers were developing various ways of amplify the fetal DNA signal to distinguish it from maternal DNA, Quake proposed a bold shortcut – skipping that step entirely. After all, he observed, the point of screening was to spot extra chromosomes in the woman's blood sample. Whether the DNA being scanned was from the fetus or from the mother didn't matter. The objective, therefore, was simply to measure the amount of DNA in fragments mapped to the different chromosomes.
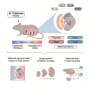
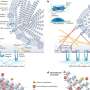
Conveniently, the cell-free DNA floating in the mother's blood circulation normally exists in short pieces, averaging 169 nucleotides in length. Quake proposed a "shotgun sequencing" strategy where he would use the very latest high-throughput gene sequencing technology and equally powerful computers to identify millions of unique sequence "tags" in the fetal DNA. Each of these tags was a 25 base-pair fragment of DNA. After the tags were identified, Quake's group could then map them to specific locations on the 23 pairs of chromosomes. By using this strategy, the researchers believed they could detect higher-than normal amounts of DNA belonging to the three chromosomes involved in the most common aneuploidies -- chromosomes 21, 18, and 13.
In the experiments reported in PNAS, Quake and his colleagues, including first author H. Christina Fan, a graduate student in bioengineering at Stanford, used their new technique to analyze DNA in blood samples from 18 pregnant women who were undergoing invasive prenatal testing (and one man, for reference).
When the amounts of DNA corresponding to each chromosome were plotted on a graph, significant, above-normal peaks appeared when a chromosome was present in three copies instead of two. These signals enabled the scientists to correctly identify the 12 women who carried aneuploid fetuses and the six whose pregnancies were normal. Their results matched those of amniocentesis or CVS test that were done on the women.
Quake said the shotgun-sequencing system is not only safer than invasive techniques but can also make a determination at an earlier gestational age, around 12 weeks.
The earlier the couple knows the status of the pregnancy, Quake observed, the better they will be prepared for whatever decision they choose to make on the basis of the information.
Quake's group is now planning a follow-up study to evaluate the test in a larger group of patients. He estimates that the cost of the test during the next phase will be about $300, with DNA sequencing accounting for the majority of the overall cost.
Source: Howard Hughes Medical Institute