Mouth bacteria can change its diet, supercomputers reveal
Bacteria inside your mouth drastically change how they act when you're diseased, according to research using supercomputers at the Texas Advanced Computing Center (TACC). Scientists say these surprising findings might lead to better ways to prevent or even reverse the gum disease periodontitis, diabetes, and Crohn's disease.
Marvin Whiteley, professor of molecular biosciences and director of the Center for Infectious Disease at The University of Texas at Austin, led the study published in April 2014 in the journal mBio.
"What we were trying to figure out," said Whiteley, "is how do these bacteria act when you're healthy, and how do they act when they're in a diseased state. The really big finding is that they do act very differently."
Bacteria share nutrients, and one species will even feed on another as they constantly interact. "The thing that we found in this paper," said Whiteley, "is that this sharing, and how they interact with each other changes quite drastically in disease than it does in health."
UT Austin researchers used shotgun metagenomic sequencing, a non-targeted way to study the all the genetic material of the bacterial communities. Whiteley and colleagues analyzed the RNA collected with the Lonestar and Stampede supercomputers at TACC. They were awarded computing allocations through the University of Texas System Research Cyberinfrastructure initiative. The research was funded by grants from the National Institutes of Health, administered by the National Institute of Dental and Craniofacial Research.
It might come as a surprise that microbes, mainly bacteria, outnumber human cells in our body by 10 to 1. And scientists have identified 10,000 different species of bacteria that live inside each person. These microbial communities are collectively known as the human microbiome. That's according to a five-year, $115 million research effort that began in 2008 by the National Institutes of Health (NIH) called the Human Microbiome Project.
"The easiest way to think of it is just the collection of bacteria that are in or on your body," Whiteley said. "We think of it as not only the bacteria, but the genetic composition. What's their DNA? And from that we can infer what these bacteria might be doing for us."
Whiteley's lab started by isolating RNA from the plaque samples collected. Study co-author Keith Turner, a postdoctoral researcher in Whiteley's lab, explained. "RNA, for those who know about computers, is kind of like the RAM (random access memory), the working memory of the cell." The RNA sample acts like a memory image or 'core dump' to reveal the processes of the as-yet unknown bacterium it came from. And unfortunately, said Turner, you can't get a full picture of the activity because there are so many molecules in the sample.
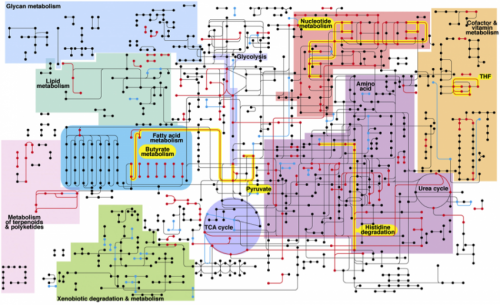
"But what you do," Turner explained, "is get what you can and profile it by sequencing, using some recent technological advances. Then it's essentially a search problem."
Turner searched a metagenomic database, essentially a vast genetic clearing house sampled from the environment instead of lab grown. He looked for matches at the NIH's Human Microbiome Project. A match told what bacterium a gene came from in the sample, and Turner tallied each match. "The more it's thinking about a certain process, the more it seems to be important to it," said Turner. "The shotgun approach, as you might imagine, is very computationally intensive, which is why we turned to TACC for some of these problems."
How big were these problems?
Turner and colleagues chose 60 different species of bacteria to represent the total community. More than 160,000 genes were analyzed, yielding 28 to 85 million reads of RNA snippets, including about 17 million mRNA reads for each sample.
His main findings show that bacteria act differently when one is healthy compared to when diseased. "The main thing that they change when they go from health to disease is that they change their metabolism," Whiteley said. In other words, a species of bacteria that ate one thing, fructose for example, can switch to a different kind of sugar to feed on if diseased.
"The kind of thing that might have taken a desktop computer a week, two weeks to run we can run at TACC in just a couple of hours," Turner said. "Stampede allows us to use 6,400 desktop computers, all at the same time. There are a lot of problems in biology that can benefit from the supercomputing approach."
Whiteley found periodontitis interesting because it's one of the most prevalent diseases on the planet. "It's an interesting disease, because the same bacteria that are in your mouth when you're healthy are the same ones, more or less when you're sick," he said.
"What our study says is that it doesn't really matter what bacteria you have, because the communities are acting very similarly," Whiteley explained. "So a healthy community has this metabolism, no matter what the members are. And a diseased community has a very different metabolism, no matter what the members are. It's this conservation of a metabolic community. "
Whiteley compared what's happening under our gums to an ecosystem in the African savannah. The interactions among 'animals' is key. "You have lions, and you have leopards, and wildebeest, and all of these animals that are there. If you look at it as a whole community, it kind of makes sense. But if you were to only take a one-acre plot out of the African savannah and look at it, it may not make sense because there may not be a lion in that one acre. So trying to understand interactions, you need to take a much larger, bigger context. And that's what this study did," Whiteley explained.
According to science results from the Human Microbiome Project, a shift to more harmful bacteria in the community is linked to wide-ranging diseases such as periodontitis, diabetes, and Crohn's disease.
Whiteley said his research can help people by helping to develop biomarkers that predict if someone's going to get sick. "Can you actually come up with a very quick way to assess the behavior of the community quickly and say, are you on the progression of moving from health to disease, and then provide some sort of preventative measure when you get there," Whiteley explained.
Pathogenic bacterial communities that rewired themselves to be harmful might also be rewired for health. It's possible in theory, anyway, according to Whiteley.
"You can manipulate bacterial populations numerically very easily. You feed them something else. So you might be able to shift them back. These are some of the ideas that we've been thinking about in our lab that might be more pervasive as we move forward."
"Medicine is going to change a lot in the next 10 to 50 years. We're going to be thinking about these sort of questions a lot more, questions like what is your microbiome actually doing, and is that impacting why you're in the doctor's office," Whiteley said.