Brown scientists map structure of DNA-doctoring protein complex
More than half of the human genome is made up of bits of mobile DNA, which can travel inside the body and insert genes into the chromosomes of target cells. This DNA doctoring not only shapes species over time, it also spreads antibiotic resistance and is used by bacteria that spread Lyme disease and by viruses linked to certain forms of cancer.
Last year in Nature, scientists working in the Brown University lab of Arthur Landy and the Harvard Medical School lab of Thomas Ellenberger announced they had solved the structure of "-integrase ("-Int), the protein "surgeon" that allows mobile DNA to cut into a chromosome, insert its own genes, and then sew the chromosome back up. That work was conducted using the lambda virus, which infects Escherichia coli (E. coli) bacteria and serves as a model that scientists use to understand mobile DNA.
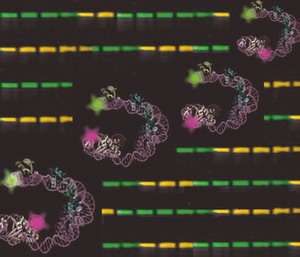
Now scientists in the Landy lab have solved the structure of a DNA-protein complex that acts as a team of "nurses," aiding "-Int during this snip-and-solder procedure known as site-specific recombination. The structure is a three-dimensional representation of the DNA within this complex. Pictured on the cover of the Nov. 17, 2006, journal Molecular Cell, it looks like DNA dressed for a party, a double helix decked with clumps of curly, colorful ribbon. By solving this structure, scientists now know how these six proteins interact with each other and fold DNA during site-specific recombination.
"Once you know how these proteins and DNA are arranged, you have a much better sense of their function," said Xingmin Sun, a postdoctoral research associate in the Landy lab and the lead author of the journal article. "And once you know their function, you begin to see how the real work inside cells gets done."
Sun said solving the structure of the DNA-protein complex called for some creativity. Because it is a string of six proteins, the complex is too big and too flexible to analyze through standard methods such as X-ray crystallography.
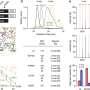
Sun used fluorescence resonance energy transfer or FRET, a technique typically used to study small protein complexes in a solution. This time, Sun used FRET to study large protein complexes in a gel. He tagged the DNA with fluorescent dyes and purified the proteins, placing them in a gel that was then shot through with light. Sun measured the wavelengths of light as they bounced between the molecules of dye. Those measurements were then fed into a special software program created by Dale Mierke, a Brown professor of medical science, which plotted their positions to create the structural map.
"The real breakthrough here is successfully using FRET to determine the structure of a large protein-DNA complex," Sun said. "Biologists now have a new tool to help them understand a variety of these complexes, including ones that control cell division, gene expression and DNA replication. So this technique represents a big advance."
Source: Brown University