Scientists crack the genome of the parasite causing trichomoniasis
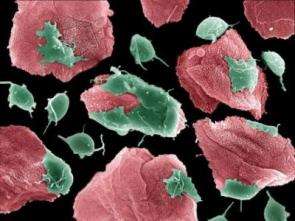
Scientists have finally deciphered the genome of the parasite causing trichomoniasis, a feat that is already providing new approaches to improve the diagnosis and treatment of this sexually transmitted disease. According to the World Health Organization trichomoniasis affects an estimated 170 million people a year and is an under-diagnosed global health problem.
Led by Jane Carlton, Ph.D., an Associate Professor in the Department of Medical Parasitology at New York University School of Medicine, the team of scientists took four years to crack the surprisingly large genome of the single-celled parasite Trichomonas vaginalis. They published the draft sequence of the parasite's genome in the Jan. 12, 2007, issue of the journal Science.
"It is a nasty bug," says Dr. Carlton. For example, in women, the parasite latches onto the vaginal lining and forms tendril-like projections into the tissue. The parasite also secretes a series of proteins that destroy vaginal epithelial cells, which make up the surface of vaginal tissue.
Women infected by the parasite can experience genital itching, vaginal discharge, and sometimes pain during urination or intercourse and an inflamed cervix. Acute infections are associated with pelvic inflammatory disease. Trichomoniasis increases susceptibility to HIV, the virus causing AIDS. Pregnant women with trichomoniasis risk delivering their babies pre-term or with a lowered birth weight. In men, trichomoniasis symptoms are usually mild such as a burning sensation after urination. The parasite can cause urogenital infections such as urethritis and prostatitis.
Worldwide, Trichomonas vaginalis infects an estimated 170 million people a year. These numbers are estimates because patients are not officially tallied. In the U.S., for example, physicians must report cases of syphilis, gonorrhea or chlamydia to local and state health authorities who report the figures to the Centers for Disease Control and Prevention. This process is not mandatory for trichomoniasis. "It is quite astonishing that this disease does not receive the attention it should," says Dr. Carlton.
The Trichomonas vaginalis genome project began in 2002 and has involved 66 researchers in 10 countries with expertise in cell biology, biochemistry and bioinformatics. Dr. Carlton performed the bulk of the work while a scientist at The Institute for Genomic Research (TIGR) in Rockville, Maryland. Her main collaborator in this project was Patricia Johnson, Ph.D., from the University of California, Los Angeles. The project was funded by the National Institute of Allergy and Infectious Diseases (NIAID), which is part of the National Institutes of Health (NIH).
Clues from the genome
There are few drugs used to treat Trichomonas vaginalis and the genome can help expand the possibilities. For example, the researchers discovered several genes and proteins not found in humans, which is important information for drug developers. One enzyme, a peptidase, is similar to a peptidase in HIV. A class of antiretroviral HIV drugs already exists to target this peptidase. "So the idea is that maybe we can use these drugs to target Trichomonas," explains Dr. Carlton.
The genome could help with diagnostics, too. The Trichomonas vaginalis genome contains a large number of repeat sequences, which could be used to devise a diagnostic test that would specifically identify this pathogen, says Dr. Carlton. The bug's large genome has around 26,000 confirmed genes, which is on par with the human genome. There may be an additional 34,000 unconfirmed genes, bringing the total gene count to around 60,000. T. vaginalis has one of the highest gene counts of any organism in the microbe, animal or plant community probably because of the puzzlingly high number of genes repeated in the genome, she says. The scientists say they still plan to work on a final gene count.
The genome tells a gory tale
The genome gives the researchers gory reading material on some of the pathogen's habits. "The genome size was the big shocker," says Dr. Carlton. "The genome was much, much bigger than we had expected, actually ten times what we had expected." All other previously sequenced parasites had much smaller genomes. "Evolutionarily speaking, having a big genome does not have many advantages if you are a parasite," she says. She and her colleagues believe the pathogen's large genome is likely to be an adaptation for its lifestyle.
Trichomonas vaginalis is typically a pear-shaped organism. When it sticks to the vaginal wall, the parasite flattens, looking almost squashed. That flattening dramatically increases its surface area. The scientists hypothesize that this trait brought the microbe a selective advantage during its evolution: A bug with a big surface area, enabled by a big genome, is better at colonizing the area it is infecting. Trichomonas also shows predatory behavior. It phagocytoses, or eats up, good bacteria in the vagina. That makes the vagina more alkaline and more hospitable toward Trichomonas and other pathogens.
When infecting women, the pathogen latches onto vaginal tissue and forms tendrils that project into the vaginal tissue. "The cytoskeleton has to change quite phenomenally in order to do this," says Dr. Carlton. The parasite also secretes a series of proteins, which destroy the vaginal epithelial cells, the cells that make up the vaginal tissue surface. For the first time, the genome project managed to identify these pathogenic virulence factors, called trichopores, she says.
Mariner elements are another novelty of the T. vaginalis genome project. They are transposons or jumping genes, which previously had only been found in animals and plants but never in protists, which are the single-celled organisms that are not plants, animals or fungi.
The genomics headache
The little bug served a sizable genomics challenge. "The big issue is that we don't really have the capability of dealing with a genome like Trichomonas," says Dr. Carlton. The sequencing technology and the computer algorithms used to assemble and align sequenced gene fragments with computers are not available to deal with this parasite. The cause of the headache for researchers: the repeats in the genome.
As Dr. Carlton explains, to sequence a genome, it is broken down into reads. These reads are snippets of DNA with 600 units or bases. Computer programs then identify similar reads, the ones with overlapping fragments of the same sequence.
These fragments are then collapsed into contiguous sequences, or contigs, so the genome is put back together like a jigsaw puzzle. Because Trichomonas has many repeating sequences, the computer algorithm got completely stuck. It could not assemble the contigs. So the scientists were stumped.
They first assured themselves their work wasn't in error and that they indeed were sequencing Trichomonas. "That made a kind of Sherlock Holmes approach necessary," she says. After all, there was little information known about the genome of this pathogen. Then bioinformatics experts and software engineers including her colleagues at the University of Maryland Steven Salzberg, Ph.D., Arthur Delcher, Ph.D. and Michael Schatz, Ph.D., had to re-work the algorithm to tackle the informatics challenge this pathogen presented. Only then could the genome project proceed to the draft now being published.
Nasty in the lab
Viewed under the microscope Trichomonas vaginalis moves quickly; it has four undulating flagella and a tail. "It is a gassy organism," says Dr. Carlton. It has special power-generating structures called hydrogenosomes. They produce hydrogen. "So it is releasing hydrogen into the liquid media, making it frothy," she says. "That is why the vaginal discharge is frothy."
The pathogen grows easily in the lab in test tubes containing some liquid media. And it has, as she says, "a real yuck factor to it." A good way to know the microbe is growing well is to smell the contents of the test tube. "It smells foul, it has a fishy odor; really nasty," says Dr. Carlton. "My technician used to get grossed out by that."
Source: New York University Medical Center and School of Medicine