Gene newly linked to inherited ALS may also play role in common dementia
Scientists at Washington University School of Medicine in St. Louis have linked a mutation in a gene known as TDP-43 to an inherited form of amyotrophic lateral sclerosis (ALS), the neurodegenerative condition often called Lou Gehrig's disease.
Researchers found the connection intriguing because studies by other groups have revealed abnormalities in the TDP-43 protein in both sporadic and inherited ALS, as well as in several other neurodegenerative disorders.
"The potential link to sporadic ALS is particularly interesting. If we can confirm TDP-43's association with inherited ALS, mutating this gene may give us a way to model sporadic ALS in laboratory animals for the first time," says senior author Nigel Cairns, Ph.D., research associate professor of neurology and pathology and immunology. "That could give us a potent tool for better understanding ALS and developing new treatments."
The study appears February 20 in Annals of Neurology. It was conducted at the Hope Center for Neurological Disorders, a partnership between the University and Hope Happens, a St. Louis-based non-profit organization dedicated to raising funds for neurological research.
Approximately 30,000 U.S. citizens have ALS, a condition that kills motor neurons, the nerve cells that control muscles. This causes gradually increasing paralysis and typically leads to death over a course of several years. Approximately five to 10 percent of all ALS cases are inherited; the rest are sporadic.
Hope Happens was founded by Christopher Hobler, a St. Louisan who developed ALS and died from the disorder in 2005. Hobler's grandfather and cousin had previously died from the disorder, and Hobler and his family founded Hope Happens to promote awareness of ALS and other neurodegenerative conditions and to raise money for research to develop new treatments and cures.
In 1993, scientists linked an inherited form of ALS to mutations in the gene for a protein called superoxide dismutase-1 (SOD1). Since then, many had thought altering the SOD1 gene's function was the most promising way to model and understand sporadic ALS.
"That has all been turned upside down in the last two years, though," says Cairns. "In that time, abnormal TDP-43 deposits have been identified in sporadic ALS cases and in some inherited forms of ALS that don't involve a SOD1 mutation."
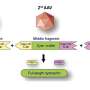
TDP-43 is an influential regulator of messenger RNA splicing, the process that edits protein-building instructions from DNA to allow the proteins to be built properly. TDP-43 abnormalities in ALS patients have included altered folding and a chemical change known as phosphorylation, both of which can radically alter the protein's function.
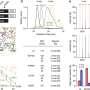
As a result, several research groups have been looking for a case where a mutation in the TDP-43 gene was linked to inherited disease. The new study is the first to tentatively establish such a link. Michael Gitcho, Ph.D., a postdoctoral research associate in Dr. Cairns' lab, and colleagues found that every member of a family affected by an inherited form of ALS had a particular mutation in TDP-43. Next, they looked at 1,505 people not related to the family and unaffected by ALS. This second search found no examples of the same mutation.
Because the family they studied is small, scientists need further evidence to confirm that the mutation is causing ALS. Researchers are working to introduce the mutated human TDP-43 gene they identified in the family into a transgenic mouse model. They hope the mouse will generate a model for ALS-like pathology.
If this affirms the link, they will begin tracing the effects of the mutation on genes whose splicing is regulated by TDP-43, working to identify key links in the chain reaction that leads to motor neuron death. These links may become new targets for pharmaceutical treatments.
What they learn may also shed light on other neurodegenerative disorders. Co-author Alison M. Goate, D. Phil., the Samuel and Mae S. Ludwig Professor of Genetics in Psychiatry, notes that abnormal TDP-43 has been found in patients with frontotemporal dementia, the second most common cause of early-onset dementia after Alzheimer's disease.
"As our understanding of these diseases progresses, we're starting to see common elements," says Goate. "This protein may allow us to link together a number of important disease entities and pinpoint new targets for therapeutic intervention."
Source: Washington University in St. Louis