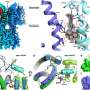
Molecular evolution of influenza A viruses circulated in Fujian Province, China
Fujian Center for Disease Control & Prevention, China, reported the molecular evolution of influenza A (H3N2) viruses in Fujian Province, south of China during the period 1996¨D2004 and demonstrated some key codons responsible for antigenic drift. The study is reported in Issue 51 of the Science in China Series C: Life Science because of its significant impact.
The FJ/411/02-like virus strains caused influenza epidemics worldwide in the 2003£04 influenza seasons. It has been shown that the viruses causing pandemics, and even year-to-year epidemics, emerge from Asia. So it is important to do surveillance and analyses of the mutation of HA1 gene of influenza virus in south of China. A/Fujian/411/2002(H3N2)-like influenza virus was recommended as one of the compositions of 2003-04 trivalent influenza vaccine for the south hemisphere and 2004-05 trivalent influenza vaccine for the north hemisphere.
Phylogenetic analysis was carried out for genes encoding hemagglutinin1 (HA1) of influenza A virus (14 new and 11 previously reported reference sequences) in this study. Phylogenetic analysis confirmed that progressive drifts occurred among our H3N2 influenza isolates over the eight flu seasons. The mutations of HA1 genes occurred from time to time, which were responsible for about four times of antigenic drift of influenza H3N2 viruses in Fujian, China.
The data demonstrated that amino acid changes were limited to some key codons at or near antibody binding sites A through E on the HA1 molecule. "The changes at the antibody binding site B or A or sialic acid receptor binding site 226 were critical for antigenic drift," noted principal investigator WenQiong Xiu, associate professor of the Department of Virology at the Fujian Center for Disease Control & Prevention, China. "The antigenic sites might change and the key codons for antigenic drift might change as influenza viruses evolve."
The study involved many experiments. The influenza strains of Fujian were directly isolated from clinical samples grown in MDCK cells or embryonated chicken eggs. Then viral RNA was extracted from the isolating fluids and was amplified by RT-PCR . Sequencing was performed and the nucleotide sequences were determined. Finally, sequence alignment and phylogenetic analysis of the sequencing data was carried out.
We also found that potential glycosylation sites were accumulated with evolution. Influenza viruses are volatile in order to survive in human body, and the viability was indeed a heritable trait of codons. The HA three-dimensional structure remains constant during antigenic drift, presumably so that the biological function of HA can be maintained. Amino acid changes in natural isolates were principally chosen from changeable substitutions that do not disrupt HA activity.
The main conclusion reported by the investigators is that it is important to monitor new H3 isolates for mutations in the positively selected codons of HA1 gene in south of Asia.
Reference: Xiu WQ, Wen YW, Shen XN, Xie JF, Yang SQ, Wu BS, Wang MA. Molecular evolution of influenza A (H3N2) viruses circulated in Fujian Province, China during the 1996-2004 period. Science in China Series C: Life Sciences 2008; 51(4): 1-8
Source: Science in China Press