A step closer to understanding, averting drug resistance
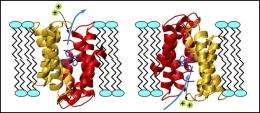
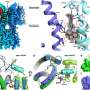
(Medical Xpress) -- The multidrug transporter EmrE functions as an asymmetric antiparallel dimer (molecule with two subunits). Drug (blue) transport from the inside to the outside of the cell membrane is accomplished by exchange between inward (left) and outward (right) facing conformations in exchange with two protons (green), directly visualized by nuclear magnetic resonance spectroscopy.
Bacterial resistance to antibiotics is growing exponentially, contibuting to an estimated 99,000 deaths from hospital-associated infections in the U.S. annually, according to the Centers for Disease Control and Prevention. One reason that this is happening is that drug resistant proteins are transporting “good” antibiotics, or inhibitors, out of the cells, leaving them to mutate.
In a paper recently published in the journal Nature, Professor of Biochemistry Dorothee Kern and collaborators including former postdoctoral student Katherine A. Henzler-Wildman, looked at how one of these drug transporters, EmrE, works. The hope is that someday a drug will be developed to impede this motion of transport.
“You have a disease and an antibiotic goes into the cells to try to kill it,” explains Kern. “But the protein EmrE takes the antibiotic and transports it out. The goal would be to find clever ways to stop EmrE from functioning as an exporter while allowing necessary nutrients to remain”
The challenge with making drugs, says Kern, is that you need to kill specific targets but nothing else.
Kern and her team studied the protein using nuclear magnetic resonance (NMR) spectroscopy, adding a drug mimetic in order to learn how the structure EmrE was designed and how it functioned as it was transporting — moving the drug from inside of the cell to the outside. Currently there are around 12 to 13 similarly known proteins within the EmrE family.
Increasing the knowledge of how this protein works will hopefully help to target all of them. Once you understand that, you could design inhibitors that do not allow the protein to transport the “good” drugs out.
“We were actually looking at a transporter in real time,” says Kern. “That is something very novel as previously these structures were only seen by X-ray crystallography, so they were frozen, and not moving.”
Kern says EmrE actually bounces back and forth between two alternate conformations, hand-shaped structures that take one molecule, pump it out, return to grab the next one, then pumps that one out (Fig. 1).
“The cool thing about discovering how the protein functions is that this was unexpected,” says Kern. “Everyone thought that the inside and the outside had two different structures.”
Kern says that she is extremely proud of her former postdoctoral student, who began working on the project at Brandeis in 2006 and has continued pushing and leading this project at Washington University in St. Louis, where she is now an assistant professor of biochemistry and molecular biophysics.
“Before Katie had worked out the kinks, my lab continued to make proteins for her and shipped them there,” says Kern. “In collaboration with postdoctoral fellow Michael Clarkson, the NMR dynamics experiments were collected at Brandeis. It’s been a real fun collaboration.”
Why are there so many resistant strains of bacteria?
Antibiotics try to kill bacteria. To avoid being killed bacteria mutate, meaning that they change the structure of the amino acids within their cell. This way the protein is no longer able to be taken over by the antibiotic.
“These bacteria survive and duplicate every 20 minutes," says Kern. “Now you have a new strain, a new form of bacteria and the designed drug no longer works. This survival and changing of composition occurs when an antibiotic encounters them.”
As more antibiotics are being used, more new strains of bacteria are created that are resistant to current those antibiotics, which is why bacteria and viruses are harder to target.
“It’s called mutagenesis—changes in the genomic material and bacteria can do this very quickly,” says Kern. “If you look at the statistics, the numbers of resistant strains are exponentially growing.”
Kern cautions against purchasing anti-bacterial products such as cutting boards, soaps and toothpaste as they are contributing to this rash of mutations.
“If you have an infection and use a high concentration of antibiotics, the bacteria are being killed before they can adapt and change,”says Kern. “If the dose is very small, they are not all getting killed, and they have time to mutate.”
Kern says this is another reason why it is so crucial to complete a prescribed course of antibiotics when you’re sick even if you’re asymptomatic —if 95 percent of the bacteria are gone, there is still five percent that can develop resistance in a matter of days.
More information: www.nature.com/nature/journal/ … ull/nature10703.html