Mother nature to the rescue
(Medical Xpress) -- Natural molecules that protect the body against disease are finding their way into the treatment of advanced cancer. Prof. Michel Revel of the Department of Molecular Genetics has played a leading role in the discovery and study of two natural molecules now employed as drugs. In the late 1970s, Prof. Revel isolated the gene for interferon-beta, a human protein that fights viral infection in the body and is used as a drug against a variety of ills, including certain types of cancer—particularly glioma and non-small-cell lung carcinoma.
Another important molecule isolated in Prof. Revel’s lab is interleukin-6, or IL-6, an immune system protein involved in defending the body against infection and inflammation. Because this protein boosts the production of blood platelets, it can offset the loss of blood cells that often accompanies intensive forms of traditional cancer therapies—chemotherapy or radiation. IL-6 may also improve blood cell formation after bone marrow transplantation.
Moreover, IL-6 can play a role in cancer treatment itself. One possibility being tested in clinical trials is to use IL-6 to improve the effectiveness of vaccines against advanced cancer such as melanoma. Studies conducted by Prof. Revel, along with the Weizmann Institute’s Prof. Lea Eisenbach, revealed that IL-6 prevents the development of metastases in animals, probably through the same immune mechanisms as those at work in vaccines. In another promising research direction, Prof. Revel’s team created an IL-6 “chimera”—a recombinant molecule that boosts IL-6’s therapeutic potential. In animal tissue, the chimera, consisting of IL-6 and its receptor fused together, has been shown to block a protein important for the survival of melanoma cells. The Weizmann scientists have started collaborating with Italian researchers to investigate the effects of the IL-6 chimera on human melanoma tumors. In addition, in collaborative research with the Institute’s Prof. Tsvee Lapidot, the IL-6 chimera molecule was found to improve the success of blood stem cell transplantation.
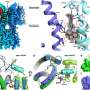
Interferons—natural anti-viral proteins—have been approved as drugs for treating viral diseases and various types of cancer. However, their use is limited by undesirable side effects; moreover, cells sometimes develop resistance to interferons or release antibodies to neutralize the drugs. These limitations may be overcome if scientists achieve a molecular understanding of how interferons bind to their cellular receptor. The laboratory of Prof. Jacob Anglister of Weizmann’s Department of Structural Biology is aiming to elucidate the three-dimensional (3D) structure of the outer part of the interferon receptor, the part that protrudes beyond the cell membrane. The scientists are also studying the complex formed by the receptor with one particular interferon—interferon-alpha2. This research is conducted using nuclear magnetic resonance (NMR) spectroscopy, a powerful tool for studying the 3D structure of proteins. The structural information gained from this study is expected to pave the way for the design of interferons and interferon-like molecules with greater therapeutic results and fewer harmful side effects.
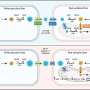
Using different technologies, Prof. Gideon Schreiber of the Department of Biological Chemistry studies the complexes formed by interferons and their receptors. Prof. Schreiber focuses on an important puzzle in the field: how is it that interferons alpha and beta bind to the same receptors on the cell membrane but produce different effects inside the cell, turning on different genes at varying intensities? Because standard methodologies have failed to produce an adequate image of the complexes’ structure, Prof. Schreiber’s team developed a new strategy. Their approach is to experimentally identify points of “docking” between the two proteins and incorporate these points into a so-called docking algorithm, a computer program that creates a 3D image of the complex. The docking points are identified using a sophisticated method, double mutant cycling, which systematically introduces mutations into the amino acids making up the protein in order to study this protein in great detail. Understanding the details of the complex interactions between different interferons and their receptors promises to provide means for designing improved interferons to fight disease.