Mutations impair childhood growth and development by disrupting organization of chromosome pairs
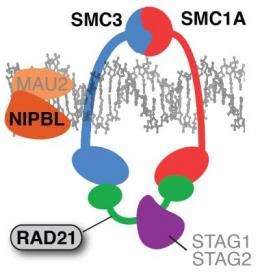
Researchers studying rare genetic disorders have uncovered insights into those diseases in biological structures that regulate chromosomes when cells divide. Focusing on the cohesin complex, a group of proteins forming a bracelet that encircles chromosome pairs, scientists have discovered mutations that disrupt cohesin, causing a recently recognized class of diseases called cohesinopathies.
"We are learning more about how these genetic abnormalities that affect cohesin play a role in human development," said study leader Matthew A. Deardorff, M.D., Ph.D., a specialist in pediatric genetics at The Children's Hospital of Philadelphia's Center for Cornelia deLange Syndrome and Related Diagnoses.
The research, carried out in children, cell cultures, and zebrafish, appeared May 24 in the American Journal of Human Genetics. Deardorff's co-study leader was Frank J. Kaiser, Ph.D., of the University of Lubeck in Germany.
The cohesin complex is already known to be involved in Cornelia deLange syndrome (CdLS), a multisystem genetic disease affecting an estimated 1 in 10,000 children. The disease has a range of severity, but classically includes mental retardation, impaired growth, heart defects, feeding problems, deformed arms and hands, and distinctive facial features.
Researchers at The Children's Hospital of Philadelphia previously were the first to discover gene mutations that cause CdLS, including forms of the disease with mental retardation and often severe limb abnormalities. The current study identified another gene, RAD21, that when mutated, causes very mild cognitive and physical impairments.
The study team first performed a genome-wide analysis of 101 children with typical CdLS and 189 children having overlapping features of the disease. None of the children had mutations in the three genes already known to cause CdLS. They identified a six-year-old boy with a deletion in a section of chromosome 8 that contains the RAD21 gene, which was known to express a cohesin protein but not previously known to cause disease. As an infant, the boy had been diagnosed with facial features similar to those of CdLS, and subsequently experienced growth retardation, but had normal cognitive development.
The researchers then focused on three additional children with deletions in RAD21 and two children with mutations within the gene, and found a similar pattern—physical features, such as short stature and distinctive facial features, overlapping with some of those seen in cohesin disorders, but with only minor cognitive delays. "These findings suggest that children who are very mildly affected may go undiagnosed," said Deardorff.
The research team did further studies in cell cultures and a zebrafish model to investigate molecular mechanisms involved in cohesin disorders. The cohesin complex includes four proteins that join in a bracelet-like structure that surrounds sister chromatids, the identical pairs that result from chromosome duplication prior to cell division. RAD21, the protein expressed by the gene with the same name, forms a clasp that closes the bracelet. A mutated RAD21 gene weakens that clasp, impairing cohesion's normal abilities to repair damage to DNA.
However, Deardorff added, the lab research does not currently explain the full sequence of molecular events, and further studies will investigate knowledge gaps in the process.
As the cost of whole-genome sequencing is rapidly dropping, Deardorff expects researchers to discover additional genes involved in cohesinopathies, offering further clues to how these diseases function in human development. "For now we can expect that patients with the RAD21 mutation will be less severely affected than those with classical CdLS," he said. "As we better understand the mechanisms of these congenital diseases, we'll continue to seek opportunities to devise more effective treatments."
More information: "RAD21 Mutations Cause a Human Cohesinopathy," American Journal of Human Genetics, published online May 24, 2012, to appear in print June 8, 2012. dx.doi.org/10.1016/j.ajhg.2012.04.019