New protein signature of breast cancer progression identified
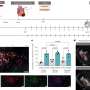
A protein signature that predicts overall survival in breast cancer patients has been uncovered in the most comprehensive survey of protein expression patterns in breast cancer progression to date.
The findings of the collaborative study led by the Max Planck Institute of Biochemistry, Germany and involving researchers in University College Dublin and the National Institute of Cellular Biotechnology, Dublin City University are published in the current issue of Cancer Research.
Breast cancer is the second leading cause of cancer death for women in the USA. Within the various subtypes of the disease, estrogen receptor negative (ER-) tumours have a relatively poor prognosis.
This study focused on identifying the proteins associated with the progression of ER- tumours and, of the 8750 proteins identified, 7800 were quantified. Proteomic analysis showed that high levels of proteins, CRABP2 and IDH2, along with low levels of SEC14L2 were predictive of overall breast cancer survival in patients.
The Irish contribution to the study, which involved Conway Fellow, Professor William Gallagher and Dr Stephen Madden (DCU), focused on the in silico validation of the candidate multi-marker signature of these 3 proteins (CRABP2, IDH2, SEC14L2).
‘As the initial discovery work by the Mann group in the Max Planck Institute was performed on a discrete set of cell culture models and clinical specimens, our work provided a key element of verification on more than 1,200 patients,”, said Professor Gallagher, Associate Professor of Cancer Biology, UCD School of Biomolecular and Biomedical Science and deputy coordinator of the SFI Molecular Therapeutics for Cancer Ireland Strategic Research Cluster. “We were also able to demonstrate that this prognostic signature retained value at the mRNA level, as the original discovery work was at the protein level”.
Commenting on the potential impact of the research, Professor Gallagher added, “The real strength of this work is that it will act as a very extensive repository for information for multiple investigators worldwide, thereby impacting on a wide range of downstream studies”.
Ideally, Gallagher and his team would hope to look at the functional relevance of the differentially expressed proteins identified in terms of their respective contributions to breast cancer and to further validate using hundreds to thousands of clinical specimens that this 3-marker signature also works at the protein level.
More information: Proteomic portrait of human breast cancer progression identifies novel prognostic markers. Tamar Geiger, Stephen F. Madden, William M. Gallagher, Juergen Cox and Matthias Mann, Cancer Research, May 1 2012 (first published online Mar 12)