Results from gut bacteria sequencing project coming in
The initial results are now coming in for a project led by the University of Colorado Boulder that is expected to eventually sequence the gut bacteria of tens of thousands of people around the world in hopes of better understanding nutrition and health.
The crowd-funded effort, known as the American Gut project, or AG, has thus far sequenced microbes from the digestive tracts of 1,589 people (as well as skin and mouth bacteria from several hundred of the participants) and has received $615,000 in donations from more than 6,700 people and four companies. Led by CU-Boulder Professor Rob Knight of the BioFrontiers Institute, the effort is the largest crowd-funded science project ever undertaken.
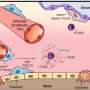
Each human is believed to harbor roughly 10 trillion microorganisms – about 10 times more than the number of cells in and on a human body – that undertake functions ranging from digesting food to strengthening immune systems. The "microbiome" includes all of the microbes and their genes interacting with the environment on and in living organisms, said Knight, who holds faculty appointments in both CU-Boulder's chemistry and biochemistry department and in the computer science department.
The AG project was founded to enable public participation in microbiome research and to expand the understanding of the variation and composition of the human microbiome, said Knight. New knowledge about the human microbiome could lead to medical interventions, the prediction and prevention of disease and the mitigation of health problems, including obesity and malnutrition.
"This is the first project where members of the public can get their microbes sequenced and the data can be a resource for anyone to use, including educators, researchers and interested members of the public," said Knight. The data gathered as part of the project is confidential in that it does not identify specific people with the exception of three participants who agreed to allow their results to be public, he said.
One preliminary new finding from American Gut study participants? Those people on the so-called Paleo Diet, which includes more protein and fiber and less carbohydrates and sodium, have fewer Proteobacteria, which have been linked to inflammation. But Paleo Diet individuals also have more of a microbe group called Firmicutes that has been associated with obesity, said Knight.
"Understanding these links between microbes and nutrition in more detail by studying more people will help us untangle the complex associations between diet, microbes and health," Knight said.
Knight and his colleagues also have compared preliminary data from the AG project with data from the Human Microbiome Project, a massive $173 million initiative funded by the National Institutes of Health that has thus far characterized the human microbiome of more than 300 healthy subjects. The results showed that oral, skin and gut microbiome assessments from both projects closely resemble each other.
The new AG data also show the effects of diet, age, exercise frequency and last antibiotic use on gut microbiomes. While antibiotic use within the past several weeks appears to have significant effects on an individual's gut microbiome, other research has shown the results depend on which antibiotic is used and which microbes were present in a person's gut prior to antibiotic use, said Knight.
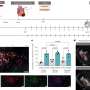
For the project, microbial DNA was isolated directly from swabs used for sampling each body site, eliminating the standard culturing step. Specific bacterial RNA genes present in the DNA were then amplified using a technique known as PCR and the genes were then sequenced with high-capacity DNA sequencers, said Knight, who also is a Howard Hughes Medical Institute Early Career Scientist.
The AG project has so far included contributions from more than 50 researchers from across the country, including graduate students, post-doctoral fellows, technicians and faculty. The Interdisciplinary Quantitative Biology program, or IQ Biology program, at CU-Boulder's BioFrontiers Institute and Department of Computer Science has been instrumental in the success of the American Gut project by supporting two members of the core team – doctoral students Daniel McDonald and Adam Robbins-Pianka.
"This support made possible the development of the infrastructure and logistics for handling tens of thousands of samples, the automated analysis of the samples and the critical biological interpretation necessary for quality of control for the American Gut project," said McDonald, who been leading the software development and data interpretation efforts for the project as part of his doctoral thesis.
"In the next phase of the study we will start looking at specific things that people do that might change the microbiome, sending out pairs of kits that could be used before and after changes in diet, exercise, drug intake and other factors," said Knight. "That will let us start answering questions about how much you can change the microbiome and in what directions these changes may occur.
"With advances in DNA sequencing, we are moving toward a world in which no infectious disease goes undiagnosed, and in which we have full knowledge of the microbes that inhabit us and our surroundings," said Knight. "By participating in this project, thousands of people are helping us make this future a reality."
For more information on the American Gut project go to americangut.org .