Genome sequencing illuminates rare Aicardi syndrome
As my inbox fills with ever more updates on the number of human genomes sequenced and the plummeting time and cost of next next next generation sequencing, I find myself hitting delete more and more often. Instead, I'm drawn to the small stories, the incremental revelations that may affect only a few individuals.
A few weeks ago, a study published in Investigative Ophthalmology and Visual Science caught my attention. Researchers at the Translational Genomics Research Institute (TGen) in Phoenix used exome and genome sequencing to probe the origins of a condition I'd never heard of — and the findings were surprising.
Aicardi Syndrome is a neurodevelopmental disorder of childhood. The three cardinal symptoms are:
- "infantile spasms." These may begin before birth and after progress to seizures
- lack of the corpus callosum, the band of nerve fibers that joins the right and left brain hemispheres
- "chorioretinal lacunae," which look like white craters in photos of the retina
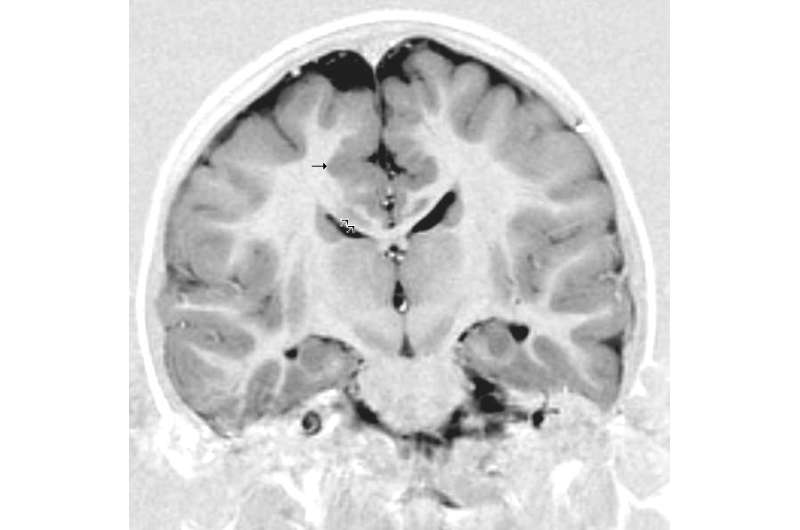
All affected children have the eye abnormality, but only some have brain manifestations. Brain imaging reveals a constellation of abnormalities, including convolutions that are too thick or too thin, cysts, enlarged spaces (ventricles), and a general asymmetry. Children tend to be developmentally delayed and intellectually disabled, with small hands, scoliosis, gastrointestinal problems, and unusual facial features.
French neurologist Jean Aicardi first described the syndrome in 1965. Only about 4,000 children worldwide are known to have it, about 900 in the U.S. Nearly all are girls; a few reports describe affected boys with XXY (Klinefelter) syndrome. They have an extra X chromosome.
The most likely explanation for the origin of Aicardi syndrome, based on whom it affects, is a dominant mutation in a gene on the X chromosome. In this rare mode of inheritance boys, lacking a second X, are much more severely affected than girls. If boys survive to be born, they may have symptoms so much more severe than those in girls that they are perhaps not even recognized as having the same syndrome.
X-linked dominant conditions are exceedingly rare, because a male can't pass it on (he'd be dead) and a female would likely be too impaired to have kids. So cases reflect a mutation that arises anew, or "de novo." The condition is genetic, but not inherited.
The TGen researchers were intrigued by reports of two boys who have normal XY male chromosomes. Might a gene not on the X cause Aicardi syndrome? The role of the X had been inferred from the preponderance of girls, not from identifying a specific responsible gene. So the investigators sequenced ten child-parent trios in search of candidate genes. Perhaps a gene on an autosome affects expression of genes on the X.
One child had a de novo mutation in a gene called TEAD1. That stands for the Tea domain, which is part of the Hippo signal transduction pathway. TEAD1 enhances the expression of several genes involved in the cell cycle and apoptosis. And it's highly expressed in the hippocampus (no link between the two hippos and the fact that it is my favorite mammal).
Further experiments showed that TEAD1 fit the bill as a candidate gene for at least some cases of Aicardi syndrome, even though it's not in databases commonly used to match mutations to phenotypes. But it is associated with an eye condition, Sveinsson's chorioretinal atrophy. Exome and genome sequencing are enabling us to connect conditions that we didn't know were connected.
The biggest surprise was that TEAD1 isn't on the X chromosome; it's on chromosome 11. And that has implications for diagnosis, which is based on clinical exam, brain imaging, and ophthalmological evidence. A handful of genetic tests are used to rule out Dandy Walker syndrome, agenesis of the corpus callosum, neuronal migrating disorders, and a few others.
Summarizes co-author Matt Huentelman, PhD, head of the Neurobehavioral Research Unit at TGen:
"Our finding suggests that the field may need to revisit how this disease is diagnosed/phenotyped. Perhaps there are male patients out there with the 'wrong' label or no label at all for their disorder simply because they weren't the 'correct' sex to receive the diagnosis of Aicardi. Or Aicardi could look similar yet different in male patients. The other big take home message for me was that for a disease that is thought to be an 'easy' one to phenotype/diagnose… we saw a wide range of phenotypic characteristics and currently no overlapping genetics."
Finding a candidate gene is a giant step forward. Not only may it refine diagnosis, but understanding the function of that gene may reveal the mechanism of the pathology—at least for some cases. And that can inspire drug discovery and perhaps other therapeutic approaches like gene or cell therapies, or even identify drugs for repurposing.
More information: "A De Novo Mutation in TEAD1 Causes Non-X-Linked Aicardi Syndrome." Invest Ophthalmol Vis Sci. 2015 Jun 1;56(6):3896-904. DOI: 10.1167/iovs.14-16261
This story is republished courtesy of PLOS Blogs: blogs.plos.org.