Volunteers help researchers sift through rare disease research literature
They refused to accept his fate. Seeking to unravel the mystery of what was making Bertrand sick, Cristina and Matthew Might found allies in the biomedical community, next-generation sequencing, the Internet and social media.
The Salt Lake City couple partnered with clinicians whose work eventually led to what they thought was the cause of Bertrand's illness: mutations in the NGLY1 gene. Through social media, the Mights found other children with similar symptoms, a key step in confirming the diagnosis.
Now they will have an army to help find a cure or treatment for the ultra-rare genetic disorder.
Mark2Cure, a project at The Scripps Research Institute's Su Lab that mobilizes crowd sourcing and computer science in the search for cures and treatments, has targeted NGLY1 deficiency for its first campaign.
Hundreds of volunteer "citizen scientists" will be recruited and trained by the La Jolla, Calif.-based grant-funded project to read and annotate abstracts of biomedical research articles in the public domain.
Working with roughly 10,000 documents, curators will identify genes, diseases and treatments in the texts that are related to NGLY1.
The goal is to attack a problem in scientific communication that can slow the search for cures and treatments: data deluge.
"One million research articles are published each year but no one is staying up-to-date with the literature as a whole," said Andrew I. Su, head of The Su Lab, a research group at Scripps, and associate professor in the Department of Molecular and Experimental Medicine.
Trained volunteers can read and annotate key terms in medical literature "at a greater scale than individual scientists could ever hope," Su said. And they can do it better than a computer.
Crowd-sourced curation of biomedical literature will allow researchers to focus on articles most useful to them and to potentially see novel links between NGLY1 and other diseases and drugs.
Such "hidden connections," Su said, could lead to re-purposing existing drugs that may help alleviate some symptoms of NGLY1 – or lead to a cure or new drug as the disease is better understood.
Mark2Cure targeted NGLY1 because of the high level of interaction between patient families and the biomedical community.
Volunteers mobilized by the NGLY1 Foundation were among 220 people who helped Mark2Cure successfully complete the beta experiment of its interactive browser-based Web application in February.
"NGLY1 is one of the newest and smallest rare diseases in the world. Our children are the ultimate underdogs," said Cristina Might, founder and executive director of the NGLY1 Foundation.
"To have an army of citizen scientists championing the NGLY1 kids and community is awe inspiring and renews one's faith in humanity."
Bertrand Might, now 7, was the first diagnosed with NGLY1. Thirty-four patients have been diagnosed since the disorder's discovery by gene sequencing in 2012; 29 patients are living with NGLY1 deficiency.
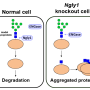
The gene NGLY1 encodes N-glycanase 1, an enzyme that recycles improperly made proteins. Without the enzyme, damaged proteins build up in cells, leading to serious health effects.
Symptoms of NGLY1 deficiency include developmental delay, movement disorders and lack of tears, among other conditions.
"Bertrand is a fighter," said Ms. Might. "First patients are usually dead patients. If we can accelerate the science, Bertrand just might live long enough to benefit from the first treatment."
Such urgency gives Mark2Cure a special sense of purpose.
"Those of us in biomedical research, we don't usually come in contact with people who could benefit from our work," Su said.
Having a real connection to patients is motivating, "and it is the same motivation that draws our volunteers – a desire to help those families and hope that we can improve their situation," said Su.
Anyone interested in contributing to this effort is encouraged to click 'start now' at http://mark2cure.org to begin the tutorials.