How bacterial and mammalian genomics interact to boost insulin resistance
The trillions of bacteria in your digestive system play a major role in your metabolism, and they're linked to your risks of type 2 diabetes, obesity and the related conditions that make up "metabolic syndrome," which has become a global health epidemic. Humans and animal models with diabetes and obesity have different gut bacteria than those who don't, and when scientists transfer microbiota from obese humans or animals to germ-free animals, the recipients are more likely to become obese or diabetic.
Now in experiments in mice reported this week in Cell Metabolism, researchers at Joslin Diabetes Centers have highlighted the ways in which the host's genes interact with the microbial genes to create such conditions, says senior author C. Ronald Kahn, M.D., Chief Academic Officer at Joslin Diabetes Center and Mary K. Iacocca Professor of Medicine at Harvard Medical School.
As a result, these researchers found that one strain of mice which were genetically prone to become obese became resistant to excess weight gain after their populations of gut microbiota were transformed simply by an sharing an environment with other mice.
These scientists also were able to identify certain bacterial strains that appear to play a positive or negative role in diabetes, obesity or related metabolic disorders, depending, in part, on the host animal's genetic makeup.
"Our hope is that if we can identify causal bacteria in these animal models, then we can look in humans for bacteria that serve the same kinds of function," says Dr. Kahn. "The goal ultimately would be to get a cocktail of purified microbes that is optimized for treatment of humans with obesity or diabetes—kind of a designer probiotic."
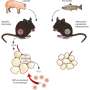
The scientists found that the three common mouse models—one prone to obesity and diabetes, one prone to obesity but not diabetes, and one resistant to both conditions—originally held very different populations of microbes in their guts. When the mice went on a high-fat diet, all of them saw dramatic change in their microbial populations. Over time, these populations became more similar among all the mice and their descendants, held in the same animal facility.
"However, when you change the microbes it has different effects on different mice, depending on the mouse's genetic background," Kahn says. "Some animals, and presumably some people, will have much more metabolic syndrome with certain microbes than other animals."
The Joslin researchers bred new generations of the three mice models and then tested whether germ-free mice who were given microbes from these three strains of mice were prone to diabetes or obesity like the donors.
Following such direct transfer of microbes, some diabetes-resistant mice gained weight and had higher glucose levels. In other animals, "even metabolically bad bacteria didn't cause a bad problem," Dr. Kahn says. "They were only a problem if the animal had the genetic susceptibility to let those bacteria grow and cause their effect."
DNA sequencing employed in the study can identify about 3,000 different bacteria in the mouse gut, of which about 300 are fairly abundant, says Dr. Kahn. Sequencing can quantify how populations of specific bacterial strains vary under given experimental conditions, allowing the investigators to look for connections with disorders in the mice.
Experiments in this field generally analyze the roles of groups of bacteria rather than individual strains. But the Joslin investigators pinpointed certain strains that correlate very strongly with conditions such as obesity and high blood glucose levels, suggesting that these strains help to cause those conditions.
The Joslin team plans to give germ-free mice some of these individual bacterial strains to see if they do help to drive changes in insulin sensitivity and other metabolic parameters. The scientists also will examine the results of altering microbiota populations in other ways, such as giving the mice antibiotics.
Lead authors on the paper were Siegfried Ussar and Olivier Bezy of Joslin and Nicholas Griffin of Washington University. Other co-authors included Shiho Fujisaka, Sara Vienberg and Samir Softic of Joslin; Luxue Deng and Lynn Bry of Brigham & Women's Hospital; and Jeffrey Gordon of Washington University. Lead support for the research came from the National Institutes of Health.