Chemists expose side effects of antimalarial drug

Malaria is a worldwide menace. According to the Centers for Disease Control and Prevention, over 500,000 individuals died from malaria in 2013 alone. While treatments for the disease exist, cures can also take a hefty physical toll. Professor of chemistry Robert Doyle in the College of Arts and Sciences, graduate students Brian Huta and Yan Nie, and an international team have begun to unravel the biochemical action of one such malarial drug.
The drug chloroquine has long been used to treat malaria, but it is not without side effects. Chloroquine kills malaria by causing the pH in certain parts of the parasite's cell to increase, preventing important biochemical reactions. Unfortunately, excessive use of the drug can be toxic to humans too. Long-term use of chloroquine can lead to side effects from gastrointestinal distress all the way up to chemical harm to the heart and death.
To better understand how chloroquine toxicity occurs, Doyle and his team set out to investigate what the drug binds to in human cells. Doyle had a hunch that chloroquine might be acting with a particular protein, saposin B, found in the lysosome of cells. Lysosomes break down and remove waste from cells, including excess lipids. People without saposin B develop a fatal, incurable condition, illustrating the importance of this protein in the cell.
"I was studying saposin B interaction with a natural lipid found in cells. It occurred to me that the lysosome is where the anti-malarial drug chloroquine also localizes after you ingest it. Since saposin B can remove damaged lipids, I wondered if it might bind chloroquine and reduce the drug's toxicity as a consequence," Doyle says.
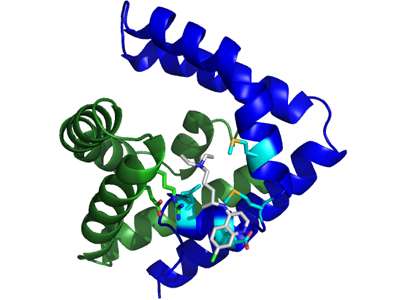
The team was pleasantly surprised to find that saposin B does bind chloroquine. This result was unexpected because saposin B is known to bind lipids—and chloroquine is not a lipid. "This is a new class of molecule for saposin B to bind," Doyle says.
Doyle, Huta and Nie confirmed the binding activity with collaborators at SUNY Potsdam. In addition, collaborators in France determined the physical structures involved in binding. "The fact that we obtained a structure of the saposin B with chloroquine bound was also wonderful, since up until now there has been no structure of saposin B with any bound molecule," Doyle says. "Quite a first, and a testament to the skills of our collaborators in France."
This new knowledge of chloroquine binding offers a suggestion as to the drug's toxicity. Because saposin B binds both lipids and the drug, prolonged use of the drug may keep saposin B busy binding chloroquine instead of removing damaged lipids from the cell, Doyle says.
Now that the team knows that saposin B binds more than just lipids, the door is wide open to investigate other conditions in which the protein may play a role. "The results suggest that saposin B might be up to a lot more than originally thought. We think that it might have a role in play in certain lysosomal storage diseases and have started a new collaboration to explore this approach," Doyle says.
The study, lead authored by Huta, was featured on the cover of the journal ChemMedChem.
More information: Brian P. Huta et al. The Lysosomal Protein Saposin B Binds Chloroquine, ChemMedChem (2016). DOI: 10.1002/cmdc.201500494