Fujitsu laboratories develops technology to accelerate analysis of genomic information
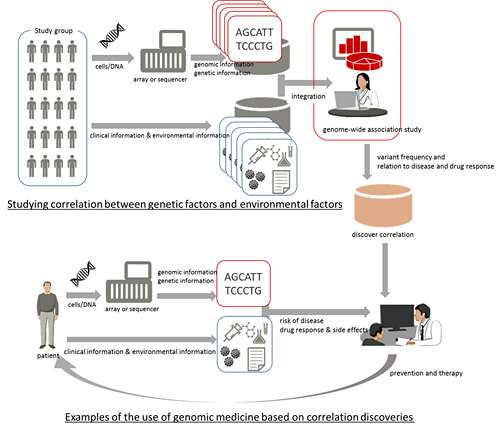
Fujitsu Laboratories announced the development of a technology that accelerates database analyses of the correlations between genomic variations and environmental information, such as disease and lifestyle habits. This technology speeds up the process by a factor of roughly 400 compared to existing methods. Thanks to advances in genomic medicine, it is possible to analyze genomic and genetic information in combination with clinical and environmental information to study the relationship between genetic factors and environmental factors. This kind of research relies on genomic information stored in databases in order to analyze the information from different perspectives, but because of the massive volumes of genomic information being handled, there is the problem of the lengthy time required for processing. Fujitsu Laboratories has greatly accelerated analysis processing by introducing a new data structure that makes it possible to rapidly analyze large-scale genomic information within a database. This technology makes it possible to acquire knowledge that previously was difficult to obtain quickly, aiding the advance of genomic medical research.
Details of this technology are being presented at the 19th International Conference on Extending Database Technology (EDBT 2016), opening March 15 in Bordeaux, France.
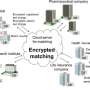
The advent of next-generation sequencers which quickly read enormous volumes of genomic information has opened up the possibility of measuring and analyzing a genome to reveal what diseases a person might be susceptible to, to predict a patient's response to a drug and the drug's side effects, and to design personalized preventative and therapeutic treatments (Figure 1, lower section). Making effective use of genomic medicine will require studying and understanding the relationship between genomic information and clinical and environmental information. With a person's entire genome being approximately three billion bases in length, there can be tens of millions of variations, known as "variants" that can account for differences between individuals. With type-2 diabetes, for example, there are dozens of variants and several lifestyle habits that are known to cause the disease, and there may be synergies among each of these factors. One method for gaining such insights is the genome-wide association study, where a huge volume of genomic information and clinical and environmental information are collected and subjected to statistical analysis (Figure 1, upper half).
Aggregating data on a single variant across a population of 100,000 people takes about one second of processing time using existing open-source database software (according to Fujitsu Laboratories' research). Accordingly, for a single disease, for example, aggregating variants at 10 million loci in a study population of 100,000 people would take roughly 120 days. Genome-wide association studies require multiple iterations of this kind of analysis, making improvements in processing speed a pressing issue.
About the Technology
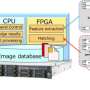
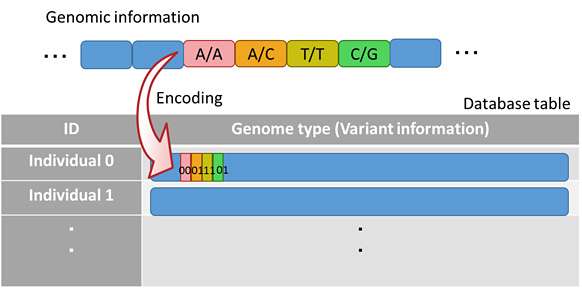
Fujitsu Laboratories has developed a data structure and its processing method for quick aggregation processing of genomic information in a database, to greatly accelerate genome-wide association studies. This structure stores an individual's genomic information in a single column in the database, and encodes information on each variant with a fixed bit length for storage (Figure 2).
This genome-type data structure has the following benefits:
1. A data structure that enables simultaneous aggregation of variants
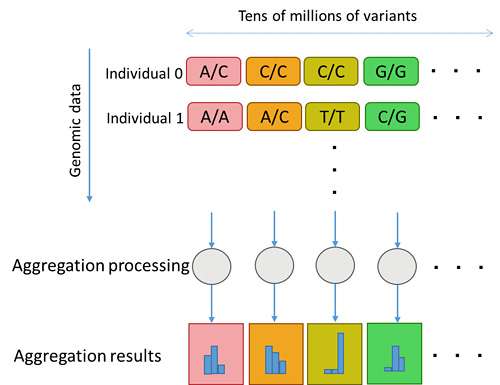
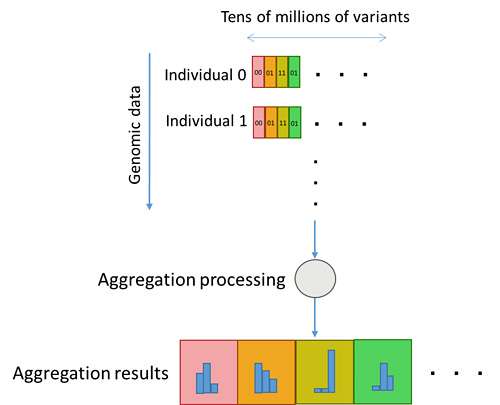
Storing each instance of variant information in a conventional database table structure required repeated database queries corresponding to the number of variants (Figure 3). With the new genome-type data structure, all variants are stored in a single column, which enables them to be aggregated simultaneously using a single query, dramatically improving the aggregation processing performance per variant (Figure 4).
2. Encoding technology allows for faster aggregation
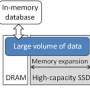
The majority of variant types can be expressed as a two-bit code using a computer. But because there are many variants that require codes of three or more bits, there is a need for variable-length data handling for codes with multiple bit lengths. When variable-length data structures are used, however, high-speed aggregation processing is no longer possible. Fujitsu Laboratories devised a method for the storing and aggregation processing of this kind of variable-length data without breaking the fixed bit-length structure, enabling high-speed aggregation processing. In addition, the encoding technique compresses the size of the genomic information to one-sixteenth of that when variants are stored as text strings. This means that data for even several hundreds of thousands of people can be handled in-memory, enabling high-speed processing.
With this technology, a genome-wide association study using all genome variants covering tens of millions of loci can be performed on a conventional computer in a short period of time. Furthermore, correlations with diseases that had been overlooked in the past due to limits on the variants studied because of time constraints can now be covered. This will help promote next-generation genomic medical research and comprehensive analyses of genomes and other molecular information in living things using "omics" big data analyses.
Fujitsu Laboratories is continuing work to further accelerate aggregation processing and to add features that will be needed for practical use. After passing through joint research with medical institutions and ethics reviews, the company plans to apply this technology to the solutions in Fujitsu Limited's Healthcare Systems Unit.