Bioinformatics software is developed to predict the effect of cancer-associated mutations
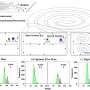
Researchers have developed free software that analyses mutations in proteins; these mutations are potential inducers of diseases such as cancer. The geneticists Asier Fullaondo and José Antonio Rodríguez, and the telecommunications engineer Gorka Prieto have created WREGEX 2.0, a versatile and fast bioinformatics application that is capable of analysing and combining the information from 40,000 proteins within the space of one minute. WREGEX 2.0 is available for the scientific community here and the journal Scientific Reports has published an article about it.
Proteins consist of chains of amino acids in which it is possible to make out short sequences with a discrete function, called "functional motifs." In some instances, these motifs have already been described; others have yet to be specified. Functional motif mutations could influence the development of diseases, including cancer. Verifying the possible changes in a protein is one of the first steps in conducting research into its function. If we bear in mind that the current draft of the human proteome is made up of over 40,000 proteins, seeking the mutations in each one is a monumental task.
Thus, the three researchers developed the bioinformatics tool. Initially, they developed a piece of software (WREGEX) to predict and automatically seek out functional motifs. They tested the programme to predict motifs that move a protein from the nucleus to the cytoplasm of a cell, the so-called nuclear exportation signals.
At the end of this research phase in 2014, they published a paper in the journal Bioinformatics. But, as José Antonio Rodríguez says, "In research, the answer to one question opens the door to more questions." The question on that occasion was: Which proteins in a sequence of amino acids could have a functional cancer-mutant motif?
The team combined the information on the sequences of all known human proteins with the COSMIC catalogue, which indexes the mutations linked to cancer. Thus appeared a new version (WREGEX 2.0) that compares normal proteins with mutants in order to predict functional motifs that have been modified and that could be linked to cancer.
"You may also have experience in how a motif functions, and you want to find out which proteins it could appear in and whether it appears modified into cancer. With this software, you can obtain candidates to start to study," explained Gorka Prieto.
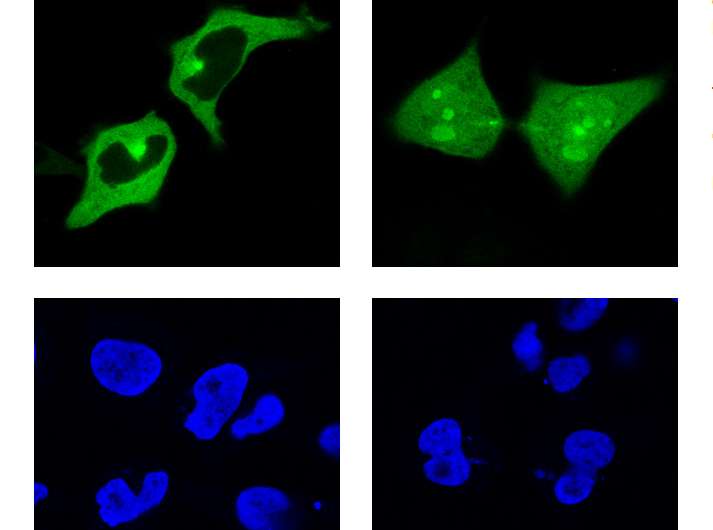
Once the bioinformatics programme had been developed, it had to be tested. To do this, they carried out a cell exportation trial. They chose various candidates that could constitute a motif responsible for moving the protein outside the cell nucleus. After modifying them according to the tumour mutations described in the COSMIC catalogue, they ran the trial again. That way, they certified that the candidates acted as an exportation signal, ensured that the mutation affected the way they worked, and that the software was therefore valid.
The tool combines three types of information: the protein sequences, the functional motifs and the cancer mutations. "One of the main features of WREGEX 2.0 is that it can simultaneously study highly complex proteomes with masses of proteins and combine information, in the case of the trial, with cancer mutations; but the door is open for using other databases containing information about other types of mutations. The advantage, moreover, is that 40,000 proteins a minute can be analysed, while with other programs, the analysis of a single protein took several minutes," explained Asier Fullaondo. With this software, it is possible to predict that the alteration in a protein may influence the development of disease, not just cancer.
So far, 13 pieces of research have already used this computing tool. Researchers in China, Japan, Korea, Germany and the United States have accessed the server. In the meantime, the researchers are already thinking about continuing with the work to improve the tool.
More information: Gorka Prieto et al. Proteome-wide search for functional motifs altered in tumors: Prediction of nuclear export signals inactivated by cancer-related mutations, Scientific Reports (2016). DOI: 10.1038/srep25869