Is Huntington's disease more common than we thought?
More people may have the potential to develop Huntington's disease than previously thought, according to a study published in the June 22, 2016, online issue of Neurology, the medical journal of the American Academy of Neurology. But the increase comes in the percentage of people who have a lower risk of developing the hereditary disease, which causes uncontrolled movements, loss of intellectual abilities, emotional problems and eventually death.
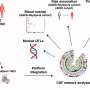
Here's how it works. The disease is passed from parent to child through a genetic mutation. The mutation is a long sequence of repeated CAG nucleotides in the huntingtin gene. The number of these repeats determines whether or not someone will develop the disease.
Everyone has two copies of the huntingtin gene—one from each parent. People who have 26 or fewer repeats on both copies of the gene will not develop the disease, nor will any of their children. People who have one copy of the gene with 40 or more repeats will develop the disease and their children will have a 50/50 chance of inheriting the gene mutation. Having between 27 and 39 repeats is known as a "gray area." People with 36 to 39 repeats have what scientists call a "reduced penetrance" of the gene. They may or may not develop symptoms of the disease.
Up until now, researchers have studied how common this reduced penetrance is mainly in people who already have symptoms of the disease and their family members. In this study, researchers use new genetic testing methods to check for the gene in the general population.
They studied the genes of 7,315 people from Canada, the United States and Scotland. Of those, 18 people had 36 or more repeats, which extrapolates to about 1 in 400 people in the general population, which is up to 10 times higher than previous estimates. Three of those people had 40 or more repeats of the gene, which is considered full penetrance. That number was consistent with previous estimates.
The study also suggests that the penetrance of the disease among people with 36 to 38 repeats is lower than previously thought, meaning that fewer people in this group would develop symptoms of the disease. For people over the age of 65, the researchers estimate that 0.2 percent of those with 37 repeats would have symptoms of the disease, compared to the 10 percent that was previously estimated. For those with 38 repeats, an estimated 2.0 percent of those over 65 would have symptoms, compared to the 19 percent previously estimated.
Study author Michael R. Hayden, MB ChB, PhD, who is a professor at the University of British Columbia in Vancouver, Canada, and also president of global research and development and chief scientific officer at Teva Pharmaceuticals, said, "It's unclear why some people with reduced penetrance genes develop the symptoms of Huntington's as early as midlife, while others reach old age with no symptoms. Additional genetic and environmental factors may modify the likelihood that a person develops the disease."
Hayden noted that while people with reduced penetrance may be at relatively low risk of developing the disease themselves, they may play a larger role in transmitting the full penetrance gene to the next generation than was previously understood.










.jpg)