Novel protein inhibitors engineered as alternative approach to potentially treat cancer
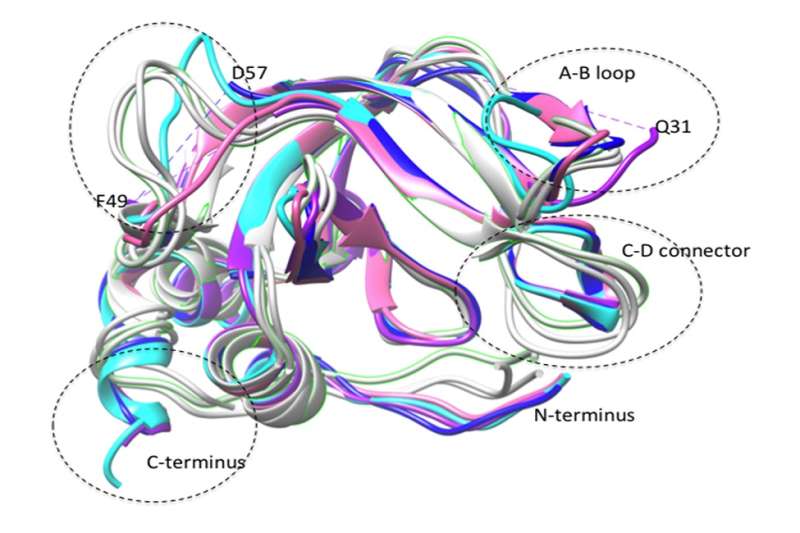
Researchers from the Charles E. Schmidt College of Medicine at Florida Atlantic University have engineered endogenous protein inhibitors of protein-degrading enzymes as an alternative approach to synthetic inhibitors for potentially treating cancer and other diseases. Results of their study, titled "Thermodynamics of Selectivity in N-TIMP/MMP Interactions," were recently published in the Journal of Biological Chemistry.
They have been investigating a group of four proteins called tissue inhibitors of metalloproteinases (TIMPs) that inhibit and control the activities of a group of 23 enzymes called matrix metalloproteinases (MMPs). Pharmaceutical companies have previously investigated the possible use of synthetic MMP inhibitors for treating cancer and other diseases, but they failed in clinical trials due to side effects, most likely because they were insufficiently specific and inhibited MMPs needed for normal physiological processes.
"There are 23 MMPs in humans that degrade or hydrolyze all of the protein components of extracellular matrices such as cartilage, bone, and basement membranes," said Keith Brew, Ph.D., a professor of biomedical science in FAU's College of Medicine. "If the regulation of their activities fails, MMPs can damage cells and extracellular structures, and their unregulated activities that are linked to serious diseases including osteoarthritis, heart disease, and cancer metastasis."
The TIMPs developed by Brew were designed to make them specific inhibitors of selected MMP targets to potentially treat osteoarthritis and cancer.
"In our study, we have been able to determine the thermodynamic profiles for the interactions of normal and mutant TIMPs with three MMPs, so that we can develop an understanding of the relationship between structural changes produced by the mutations and the changes in their abilities to inhibit different MMPs," said Brew.
To accomplish this, the researchers used a technique called "isothermal titration calorimetry" (ITC), which measures the minute amounts of heat that are given out or absorbed when two molecules interact together (the enthalpy of binding). When they measured how this varies as they increased the quantities of one protein under different conditions of temperature and solvent, they were able to determine the enthalpy change (heat absorbed or released), entropy change (a measure of dynamics in the proteins and their surroundings) and heat capacity change, enabling them to separately estimate the changes in protein and solvent dynamics.
Unlike most other protein-to-protein interactions, they found that TIMP/MMP interactions are driven by entropy increases, and not by enthalpy (which derives from specific interactions between chemical groups in the interacting molecules). They found that proportions of entropy coming from internal dynamics in the protein and from the solvent varies for different interactions, and is affected by TIMP mutations, but that the MMP component has a major role in determining the proportions.
"When we think of two proteins binding together, we have to consider that they are dynamic molecules, each consisting of a population of interconverting three-dimensional structures," said Brew. "Complex formation involves the selection of subpopulations that are compatible with each other in structures and dynamics, a process called 'conformer selection.' And, it appears that the MMP component has a major role in this process making the rational design of selective mutants difficult."