Discovery of new strains of the HTLV-4 virus in hunters bitten by gorillas in Gabon
Scientists from the Institut Pasteur and the CNRS have identified two new strains of the HTLV-4 virus in two hunters who were bitten by gorillas in Gabon. These findings, published in the journal Clinical Infectious Diseases, support the notion that gorillas represent a major source of infectious agents that can be passed on to humans.
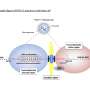
Many of the viral pathogenic agents that have emerged in humans in recent decades are of animal origin - including SARS coronavirus, avian influenza virus, hantaviruses, Ebola virus, Marburg virus and Nipah virus. After the initial contact between species, some of these viruses used a variety of evolutionary mechanisms to adapt to their new human host. Scientists from the unité d'Epidémiologie et physiopathologie des virus oncogènes (Institut Pasteur/CNRS), directed by Antoine Gessain, are working on a group of RNA viruses known as HTLV retroviruses. In 2 to 8% of cases, HTLV type 1 results in severe forms of leukemia/lymphoma or neuromyelopathy.
There are four types of HTLV virus (types 1 to 4). The animal reservoir for all four types is non-human primates, especially gorillas and chimpanzees. These viruses can be spread by sexual contact, blood transfusion or breastfeeding. Nearly 20 million people worldwide are thought to be infected by HTLV-1, especially in Japan, the Caribbean, Latin America and tropical Africa. HTLV-2 mainly affects Native American populations, some Pygmy populations and intravenous drug users; approximately a million people are infected worldwide. The HTLV-3 virus has only been observed in a small number of people in Cameroon living in close contact with infected non-human primates. HTLV-4 had previously only been identified in one person living in southern Cameroon, and the origin of the infection was unable to be traced.
Scientists from the Institut Pasteur and the CNRS, working in cooperation with the International Center for Medical Research in Franceville, Gabon (the CIRMF) and the Pasteur Center in Cameroon, set about identifying HTLV retroviruses in people at risk of contact with non-human primates. By using molecular screening to examine blood samples from 300 people who had been bitten by monkeys in Central Africa, they were able to identify the HTLV-4 virus - previously only observed in Cameroon - in two individuals in Gabon.
These individuals reported having been severely bitten by a gorilla during hunting activities, which seems to confirm the zoonotic origin of the virus. The fact that this virus was only found in two people who had been bitten by a gorilla and not in anyone who had been bitten by a chimpanzee or a smaller monkey suggests that the virus is specifically transmitted by gorillas. Moreover, the gorilla bites occurred several years before the blood samples were taken, revealing the chronic persistence of this virus in humans.
The scientists also discovered that one of the two new strains is divergent from the previously known strain of HTLV-4, highlighting the genetic diversity of this human virus.
This research supports the idea that gorillas should be seen as key reservoirs for infectious agents that can be passed on to humans. The findings suggest that there are probably several other cases in Cameroon and Gabon, as well as in neighboring countries. Work now needs to be done to identify diseases that may be caused by the HTLV-3 and HTLV-4 retroviruses.
More information: Léa Richard et al, Zoonotic Transmission of Two New Strains of Human T-lymphotropic Virus Type 4 in Hunters Bitten by a Gorilla in Central Africa, Clinical Infectious Diseases (2016). DOI: 10.1093/cid/ciw389