Variation in 'junk' DNA leads to trouble
All humans are 99.9 percent identical, genetically speaking. But that tiny 0.1 percent variation has big consequences, influencing the color of your eyes, the span of your hips, your risk of getting sick and in some ways even your earning potential.
Although variants are scattered throughout the genome, scientists have largely ignored the stretches of repetitive genetic code once dismissively known as "junk" DNA in their search for differences that influence human health and disease.
A new study shows that variation in these overlooked repetitive regions may also affect human health. These regions can affect the stability of the genome and the proper function of the chromosomes that package genetic material, leading to an increased risk of cancer, birth defects and infertility. The results appear online in the journal Genome Research.
"Variation is not only important for how genes and proteins function, but it can also occur in the noncoding, repetitive portions of the genome," said Beth A. Sullivan, Ph.D., senior author of the study and associate professor of molecular biology and microbiology at Duke University School of Medicine.
"What we found in this study is probably the tip of the iceberg," Sullivan said. "There could be all sorts of functional consequences to having variation within the complex, repetitive portion of the genome that we don't know about yet."
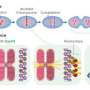
Even though the sequence of the human genome was declared complete more than a decade ago, it retains several glaring gaps, especially in the repetitive sequences around centromeres, the twisty ties that hold a pair of chromosomes together in a floppy X shape and coordinate their movement during cell division.
These centromere sequences—called satellite DNA—are made up of blocks of exactly 171 A's, C's, T's and G's, repeated over and over for millions of base-pairs. Researchers once believed that each chromosome contained a single stretch of this satellite DNA, which determined where its centromere would reside. But a few years ago, Sullivan's lab discovered that many human chromosomes possessed more than one of these regions, and depending on the individual, the centromere could form at either site.
In this study, Sullivan wanted to see how the chromosome decides where to put its centromere, and whether one site builds a "better" centromere than the other. Of the 23 pairs of human chromosomes, she focused on chromosome 17, which is structurally rearranged or mutated in many different cancers and birth defects.
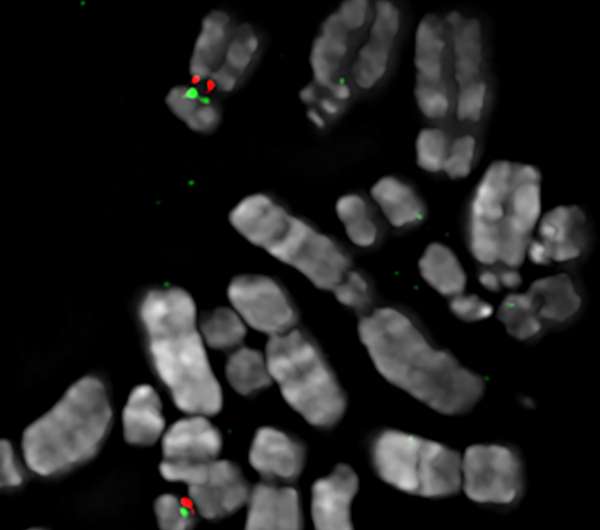
First, Sullivan and her team combined molecular and visual assays, stretching the chromosome out into long chromatin fibers that were painted with fluorescent probes to map the variation in genomic sequence at the two different regions of satellite DNA. Then they looked at each satellite region for the presence of proteins necessary to construct a fully functioning centromere.
The researchers found that genomic variation at one of these satellite DNA regions—either in the size or sequence of its repeated 171 base pair units—ultimately determines whether the centromere is built at the primary site or the alternate site.
When they interrogated samples from a human DNA bank, they found that about 70 percent of humans have little genomic variation at the primary site, while 30 percent have differing degrees of variation. Most of the time, the centromeres aren't built at the primary site if it contains variation and instead are assembled at the "backup" site nearby. But when this happens, the result may be a dysfunctional centromere that is architecturally unsound and an unstable chromosome that may be present in too many or too few copies.
"It is immensely fascinating to think that there are so many people walking around who are essentially centromere mosaics," said Sullivan. "One of their centromeres, on one of their chromosomes, has the potential to be dangerously unstable, and it could affect their ability to reproduce, or predispose them to cancer."
In the future, Sullivan plans to investigate just how big of a risk the variant satellite regions pose for those who carry them, and possibly develop a way to use these sequences as biomarkers for the chromosomal defects that can lead to disease.
More information: Genome Research, DOI: 10.1101/gr.206706.116