Team examines the molecular basis of brain plasticity and the manner in which neurons 'learn'
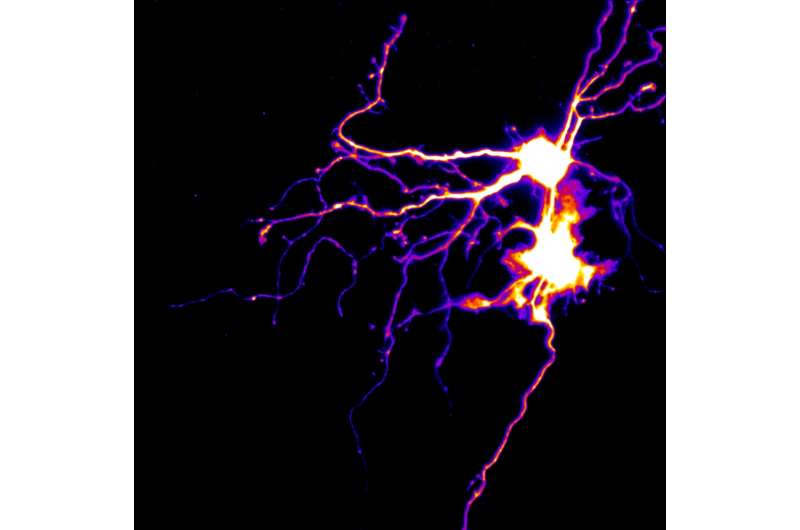
UC Santa Barbara neuroscientist Kenneth S. Kosik has been studying the brain for decades. His UC Santa Barbara neurobiology lab focuses on the evolution of synapses that connect neurons and the genetics of Alzheimer's disease. In particular, Kosik's team is interested in the underlying molecular basis of plasticity and how protein translation at synapses affect learning.
In a new paper published in the journal Neuron, Kosik explores the nature of brain plasticity and proposes a theory about how neurons learn.
"It's fairly well established scientifically that the learning units in the brain are the synapses," said Kosik, UCSB's Harriman Professor of Neuroscience and co-director of the campus's Neuroscience Research Institute. "Many neuroscientists think that synaptic learning requires new proteins made locally right at the synapse, which acts as its own control center."
A neuron are large cell that contains a cell body and dendrites, branched extensions along which impulses received from other cells at the synapses are transmitted to the cell body. In theory, learning takes place at synaptic junctions. But the sheer number of synapses—multiple thousands—makes it unlikely that all of them can make the RNA responsible for creating new proteins.
So Kosik did the math. Using available techniques, he computed the actual copy numbers of RNA in cells, which are derived from two copies of DNA in a chromosome.
"Dendrites don't actually have many RNAs, but they obviously have enough because they get the job done," Kosik explained. "What is a surprise is that they do it with a relative paucity of RNAs. That is, there are many synapses beyond the reach of any RNA and therefore those synapses are not accessible to plasticity. If you have a large portion of the brain that can't engage in learning, then what's going on here?"
Kosik noted that having a relatively small number of RNAs allows synapses to leverage increased dynamic range. The best analogy is an audio system that functions with sound fidelity at low and high volumes. Dendrites need to function with fidelity when their inputs are few or many. Having a small number of RNAs provides a quantitative space to enlarge the pool dynamically when traffic into the dendrite is high.
"The dynamic range allows dendrites to double or triple or even quadruple their learning capacity in accordance with the amount of information coming in," Kosik explained. "It also allows for sparse coding."
Another concept in neurobiology, sparse coding plays a role in how neurons process incoming information by using the representation of our memories and perceptions through the strong activation of a relatively small set of neurons. For each stimulus, this is a different activated subset utilized from the large pool of all available neurons.
Kosik explained the concept in terms of discerning odors. Too many odors exist for each one to have a unique pattern of firing neurons. Rather, the brain creates small maps. One odor might have 10 neurons that encode it, seven of which also encode a different odor, creating an overlap.
"In the same way, this idea of the dendrites having a relatively few number of RNAs allows them to receive a fewer number of inputs," Kosik said. "There are lot of inputs coming through the dendritic tree, but only a few of them are capable of learning. We call the type of learning that goes on in neurons plasticity. So the dendrite only learns from impulses where protein synthesis is available—an example of sparse coding.
"The RNAs near synapses inform us as to those synapses which have undergone plasticity, but like learning itself, the RNAs are not static," Kosik added. "The RNAs like where they are. They do a good job where they are, but they eventually degrade and have to be remade, and in so doing, they may not necessarily return to their original location. Those small changes may impair our access to a memory, but now another nearby synapse is open to novelty and has an opportunity to learn a new thing."
More information: Kenneth S. Kosik. Life at Low Copy Number: How Dendrites Manage with So Few mRNAs, Neuron (2016). DOI: 10.1016/j.neuron.2016.11.002