Hi-res view of protein complex shows how it breaks up protein tangles
Misfolded proteins are the culprits behind amyotrophic lateral sclerosis (ALS), Alzheimer's disease, Parkinson's disease, and other neurodegenerative brain disorders. These distorted proteins are unable to perform their normal functions and cause devastating problems for neurons.
Currently, there is no way to untangle the knotted mass of these proteins to treat disease.
A high-resolution view of the structure of Hsp104 (heat shock protein 104), a natural yeast protein nanomachine with six subunits, has been acquired by James Shorter, PhD, an associate professor of Biochemistry and Biophysics in the Perelman School of Medicine at the University of Pennsylvania, and colleagues at the University of Michigan. The Shorter lab has been working on Hsp104 for close to a decade as a way to dismantle harmful protein clumps in disease. The team described their findings in Science this week.
Shorter teamed with colleagues at Michigan who use cutting-edge cryo-EM to provide the clearest image to date of Hsp104 in action. The Penn team provided the highly purified Hsp104 proteins for the study.
"This superb collaboration has yielded the highest resolution picture of Hsp104 caught in the act of processing a protein," Shorter said. "We can now see the moving parts of the Hsp104 complex and how we might tune it to optimally attack neurodegenerative disease proteins."
Hsp104 pulls in proteins it "processes" through a central channel, but scientists had not seen this at high resolution before this study. "With this more-in-focus view, we can see parts of its structure that we want to engineer to make better on-target therapeutics for neurodegenerative diseases," said JiaBei Lin, PhD, co-author and postdoctoral fellow in the Shorter lab.
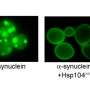
Normally, Hsp104 is a "disaggregase" enyzme, which dissolves previously aggregated proteins and helps them acquire the correct shape. Although Hsp104 is found in most organisms on the planet, it has no analogue in humans or animals. Shorter asked whether it could be introduced as a drug to dismantle the protein clumps that characterize some diseases. In previous studies, Shorter's lab established that the natural version of Hsp104 is active against neurodegenerative proteins such as alpha-synuclein.
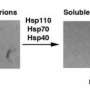
Hsp104 pulls out one polypeptide at a time from the tangles of protein fibrils. The six subunits of the Hsp104 complex hydrolyze ATP as it climbs up the polypeptide strand, which ultimately gets pulled out of the aggregate. Once released, the polypeptide can refold or be degraded.
The team has already made some tweaks to Hsp104 by mutating specific residues to enhance its activity. Working to break up TDP-43, FUS, and alpha-synuclein disease clumps, the reprogrammed Hsp104 pulls these proteins apart better.
"It appears to pull substrates through stepwise, like a ratchet," said senior study author Daniel Southworth, PhD, an assistant professor at the University of Michigan Life Sciences Institute. "We can see how the proteins in the machine rearrange between different states to grab onto the next site on the substrate."
"The study helps us to understand how cells can break apart toxic protein aggregates to restore protein function," Shorter said. "Finally having a clear picture of this remarkable nanomachine will empower our designs for therapeutic versions that work in humans."