Team presents an online tool to extract drug toxicity information from text
There is an increasing interest in more sophisticated search engines that are tailored to cope with the complexity of biomedical data, not only enabling more targeted search queries but also easier integration and construction of biological knowledge bases and analysis of experimental datasets.
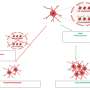
The LimTox online software tool integrates state-of-the-art text mining, machine learning and language technology methods in order to empower its biomedical semantic search engine. LimTox allows retrieval and ranking of chemical and biological entities, interactions between them, visualization of chemical structures of compounds detected automatically in the text, and generation of entity relation network graphs.
The online tool provides information on drug hepatotoxicity extracted from abstracts and full text papers from the biomedical archive PubMed, the European Public Assessment Reports (EPAR), published by the European Medicines Agency (EMA), and the United States New Drug Application (NDA).
The LimTox webserver can help researchers and clinicians to retrieve associations to adverse reactions by using simple keyword searches and queries particularly optimized to handle entities such as chemicals and genes. It's free and open to all users at http://limtox.bioinfo.cnio.es/
"There has already been some substantial work on text mining of genes, but far less on chemicals," explains Martin Krallinger, head of the Biological Text Mining Unit and main author of the paper. "To address this limitation, we have implemented this system."
A systematic strategy for efficient online access to biological and chemical information contained in scientific literature and medical agency reports is critical for scientific intelligence and decision making in areas such as chemical biology, drug discovery, toxicology and pharmacogenetics.
LimTox has a special focus on adverse reactions and chemical compound toxicity with emphasis on drug-induced liver injury, including substances that cause worsening of hepatic function and hepatocarcinogenesis. It also enables systematic access of information related to other adverse reactions (nephrotoxicity, cardiotoxicity, thyrotoxicity, phospholipidosis), alteration of biochemical liver markers and key enzymes for drug metabolism (P450 cytochromes -CYPs).
"Among the potential candidate toxicological end points, hepatotoxicity represents one of the most critical toxic effects at the organ level. The liver is a fundamental organ examined in toxicology studies due to its central role in metabolic, excretory and synthetic biochemical pathways, and the mechanisms leading to drug-induced liver toxicity are particularly complicated," says Krallinger.
More information: Andres Cañada et al, LimTox: a web tool for applied text mining of adverse event and toxicity associations of compounds, drugs and genes, Nucleic Acids Research (2017). DOI: 10.1093/nar/gkx462