Three Klebsiella species cause life-threatening infections and share drug resistance genes
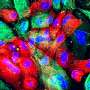
A team of US researchers has discovered that three different species of Klebsiella bacteria can cause life-threatening infections in hospital patients and that all three share genes that confer resistance to the most commonly used antibiotics. The study, published this week in mSphere, an open-access journal of the American Society for Microbiology, improves physicians' understanding of Klebsiella infections and could point toward better ways to fight multi-drug resistant strains of these bacteria.
"Since 2001, we've seen a global explosion of drug-resistant Klebsiella infections," says S. Wesley Long, Associate Medical Director of the Diagnostic Microbiology Lab at Houston Methodist Hospital in Texas and lead author of the study. "They are drug-resistant bacteria that are increasingly difficult to treat because they are resistant to many of the available antibiotics."
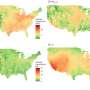
Klebsiella are a type of bacteria that cause healthcare-associated infections, which can take the form of pneumonia, sepsis, wound infections and urinary tract infections. Healthcare-associated infections numbered more than 700,000 in the US in 2011 and up to 50 percent of invasive , multidrug-resistant K. pneumoniae infections have been fatal in some studies. In the last two decades, antibiotic-resistant Klebsiella infections have been on the rise around the world.
Long and his team wanted to investigate the nature of Klebsiella infections by studying a large, comprehensive, population-based sample collection. "We need to understand the pathogen on a population level, then we can use the bacterial genomes to predict virulence or antibiotic resistance of the strain, or mortality," notes Long, a clinical microbiologist.
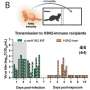
In a study previously published in mBio, the researchers sequenced the entire genome of 1,777 Klebsiella from clinical specimens across the greater Houston area. Until now, Klebsiella pneumoniae was thought to be the culprit in most serious Klebsiella infections. However, the research team noticed a group of 28 samples that looked genetically different.
"We built a genetic family tree, essentially, and 28 strains stick off the tree as outliers. These are cousins five times removed and we wondered what are these guys doing at the family reunion?" says Long.
It turned out that, depending on the collection, between 2-12 percent of the samples had been misidentified as Klebsiella pneumoniae, and were in fact two related species, Klebsiella variicola or Klebsiella quasipneumoniae. K. variicola and K. quasipneumoniae had previously been characterized as commensal, nonpathogenic bacteria of the GI tract or agricultural pests, which rarely caused human infections. Long's team found they were capable of causing invasive and severe infections in patients, with the same rate of mortality as K. pneumoniae.
"Not only are these cousins of K. pneumoniae causing similar infections, but they are also sharing these powerful drug resistance genes," says Long. The sequencing of all bacterial genetic material present showed that all three Klebsiella species were sharing drug-resistance genes amongst themselves—including at least two genes that code for powerful enzymes that disable a broad spectrum of penicillin-like antibiotics.
Long says that these findings will not likely change the way Klebsiella infections are treated. "But in the race of trying to understand pathogens and find new antibiotics, or therapies outside the box of traditional antibiotics, this expands our knowledge of what pathogenic Klebsiella look like." Other genetic traits that the three Klebsiella species share might be exploited as an Achilles' heel weakness and attacked by new, targeted therapies. Long also says the work stresses the importance of doing large, comprehensive, population-based studies that take a close genetic look at patient samples: "If you are not looking, you don't know what you're missing."