How the immune system identifies invading bacteria
The body's homeland security unit is more thorough than any airport checkpoint. For the first time, scientists have witnessed a mouse immune system protein frisking a snippet of an invading bacterium. The inspection is far more extensive than researchers imagined: the immune system protein, similar to those in humans, scans the bacterial protein in six different ways, ensuring correct identification.
"This was very surprising," says Howard Hughes Medical Institute (HHMI) Investigator Eva Nogales, a structural biologist at the University of California, Berkeley. "The immune system protein uses many protein parts, including some of previously unknown function."
This discovery, reported November 16 in Science, reveals details of a fundamental process the immune system uses to recognize pathogens that have gained entry into cells. The work also helps explain why it's hard for certain bacteria—such as the human pathogens Salmonella, Pseudomonas, and Legionella—to evade immune system detection.
A multidisciplinary, international effort allowed scientists to witness this pathogen-detection system firsthand. HHMI Investigator Russell Vance had been studying the NLR superfamily of immune system proteins, which plants and animals use to detect pathogens that have slipped inside cells. He wanted to see one such protein, called NAIP5, as it inspected bits of protein shed by the disease-causing bacterium Legionella pneumophila. Earlier genetic studies had identified NAIP5 as an important player in host resistance to Legionella, and Vance's team wanted to take a closer look. So a student in his lab, Jeannette Tenthorey, teamed up with a student in Nogales's lab, Nicole Haloupek, who used a state-of-the-art imaging technique called cryo-electron microscopy (cryo-EM) to visualize the proteins.
With cryo-EM, scientists mix up a solution of proteins, freeze it, and then blast it with a beam of electrons. The electrons scatter as they hit the proteins, and then pass through a lens to a detector. From the resulting images, researchers can construct detailed three-dimensional protein structures.
Other scientists had previously tried imaging the NAIP5 protein as it scrutinized bits of bacteria. But the images lacked important details about which parts of the protein touched the bacteria. To skirt this problem, Nogales and Vance tapped into the computer modeling expertise of researchers at the Rocasolano Physical Chemistry Institute in Madrid.
"Seeing these proteins self-assemble was really quite beautiful and fascinating," Nogales says.
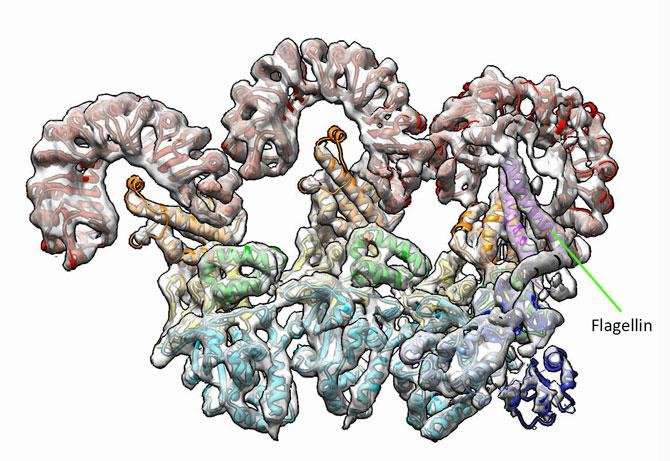
The researchers discovered that NAIP5 performs an in-depth inspection of bits of the bacteria's flagella, the tail-like appendages that many disease-causing bacteria use for locomotion. "This is a very effective immune response," says Vance, a microbiologist and immunologist, also at UC Berkeley. "It helps us understand why the pathogen can't escape just by mutating."
Bacteria can't simply hide from the immune system by making minor changes to flagella proteins, he explains. And larger changes that might let bacteria evade detection could meddle with locomotion.
The team tested the idea by creating mutant strains of Legionella and introducing them to the immune system proteins. Sure enough, minor mutations to a bacterial flagella protein weren't enough to trick NAIP5. But more significant mutations interfered with the flagella so much that the bacterium had trouble moving around.
Intensive frisking by the immune system suggests that it is careful to identify a threat before taking out the big guns, Vance says. After glomming onto the bacterial protein snippet, the immune system protein recruits a second protein, forming a complex called an inflammasome. The second protein then sounds an alarm that the cell has been invaded, triggering events that culminate in a dramatic form of cell death.
"The cell literally bursts open," Vance says. This dramatic finale—called pyroptosis—is a good thing if a bacterium is trying to take up residence in a cell, he says, but the chain of events can provoke disease if it happens too often. That's why it's important that the immune system is thorough, and the response is highly specific to the bacterium's flagella, he says.
More information: J.L. Tenthorey el al., "The structural basis of flagellin detection by NAIP5: A strategy to limit pathogen immune evasion," Science (2017). science.sciencemag.org/cgi/doi … 1126/science.aao1140