Mutant gene network in colon cancer identified
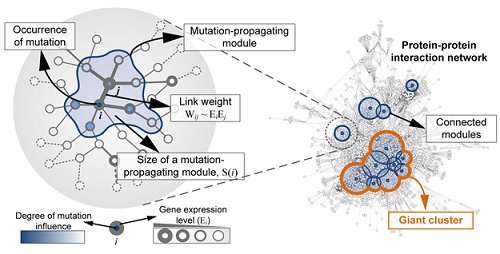
The principles of the gene network for colon tumorigenesis have been identified by a KAIST research team. The principles will be used to find the molecular target for effective anti-cancer drugs in the future. Further, this research gained attention for using a systems biology approach, which is an integrated research area of IT and BT.
The KAIST research team led by Professor Kwang-Hyun Cho for the Department of Bio and Brain Engineering succeeded in the identification. Conducted by Dr. Dongkwan Shin and student researchers Jonghoon Lee and Jeong-Ryeol Gong, the research was published in Nature Communications online on November 2.
Human cancer is caused by genetic mutations. The frequency of the mutations differs by the type of cancer; for example, only around 10 mutations are found in leukemia and childhood cancer, but an average of 50 mutations are found in adult solid cancers and even hundreds of mutations are found in cancers due to external factors, such as with lung cancer.
Cancer researchers around the world are working to identify frequently found genetic mutations in patients, and in turn identify important cancer-inducing genes (called 'driver genes') to develop targets for anti-cancer drugs. However, gene mutations not only affect their own functions but also affect other genes through interactions. Therefore, there are limitations in current treatments targeting a few cancer-inducing genes without further knowledge on gene networks, hence current drugs are only effective in a few patients and often induce drug resistance.
Professor Cho's team used large-scale genomic data from cancer patients to construct a mathematical model on the cooperative effects of multiple genetic mutations found in gene interaction networks. The basis of the model construction was The Cancer Genome Atlas (TCGA) presented at the International Cancer Genome Consortium. The team successfully quantified the effects of mutations in gene networks to group colon cancer patients by clinical characteristics.
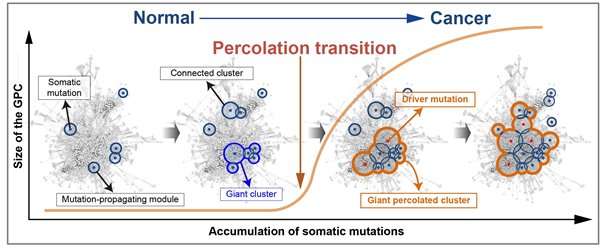
Further, the critical transition phenomenon that occurs in tumorigenesis was identified using large-scale computer simulation analysis, which was the first hidden gene network principle to be identified. Critical transition is the phenomenon in which the state of matter is suddenly changed through phase transition. It was not possible to identify the presence of transition phenomenon in the past, as it was difficult to track the sequence of gene mutations during tumorigenesis.
The research team used a systems biology-based research method to find that colon cancer tumorigenesis shows a critical transition phenomenon if the known driver gene mutations follow sequentially. Using the developed mathematical model, it can be possible to develop a new anti-cancer targeting drug that most effectively inhibits the effects of many gene mutations found in cancer patients. In particular, not only driver genes, but also other passenger genes affected by the gene mutations, could be evaluated to find the most effective drug targets.
Professor Cho said, "Little was known about the contribution of many gene mutations during tumorigenesis." He continued, "In this research, a systems biology approach identified the principle of gene networks for the first time to suggest the possibility of anti-cancer drug target identification from a new perspective."
More information: Dongkwan Shin et al, Percolation transition of cooperative mutational effects in colorectal tumorigenesis, Nature Communications (2017). DOI: 10.1038/s41467-017-01171-6