March 9, 2018 report
Study suggests some CpGs in the genome can be hemimethylated by design

A pair of researchers at Emory University has found that some CpGs in the genome can be hemimethylated by design, rather than by chance. In their paper published in the journal Science, Chenhuan Xu and Victor Corces describe their study of DNA methylation and the fate of hemimethylated DNA in daughter strands after replication. Jafar Sharif and Haruhiko Koseki with the Developmental Genetics Group, Center for Integrative Medical Sciences in Japan offer a Perspective piece on the work done by the team in the same journal issue.
DNA methylation (when methyl groups are added to the DNA molecule) is a modification that serves to regulate gene transcription, embryonic development, and cell differentiation in plants and animals. In animals, specifically mammals, methylation happens at the CpG dinucleotides symmetrically, which results in corresponding cytosine residue on the CpG components. But, as the researchers note, this process is halted during replication, a period during which a daughter strand that is unmethylated and a methylated parent strand work in conjunction to create a methylated CpG dyad. This is called hemimethylated DNA.
Prior research has shown that such hemimethylated DNA generally tends to fully methylate, or in some cases, to unmethylate by dilution. But approximately 10 percent of trophoblast or embryonic stems cells remain hemimethylated. Until now, it has been a mystery whether this occurs by simple chance or if there is some other process at work. In this new effort, the researchers discovered that at least some of these instances are due to design. Furthermore, they found that hemimethylated instances are inherited and happen over the course of several cell divisions.
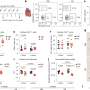
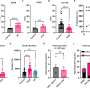
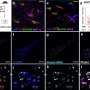
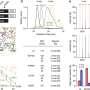
To learn more about the process, the team mapped the DNA methylome that was targeted by three types of DNA methyltransferases, which showed some of the interactions that occur between DNMIs and daughter cytosines. They also showed inheritance of hemiCpGs at binding sites in pluripotent cells. More specifically, they found that during replication, at the fork, hemimethylated DNA is held by UHRF1, a reader protein, and thereafter, DMNT1 was recruited, which served to convert CpGs to symmetrical methylation. This resulted in reinstating the original methylation symmetrical pattern that existed prior to replication. They found also that DNMT1-bound emerging DNA fragments were overwhelmingly hemimethylated. They that it was elimination of the cause of hemimethylation factors that resulted in the frequency of chromatin interactions, which, they claim, suggests that hemimethylation might actually serve as means for stabilizing chromatin interactions.
More information: Chenhuan Xu et al. Nascent DNA methylome mapping reveals inheritance of hemimethylation at CTCF/cohesin sites, Science (2018). DOI: 10.1126/science.aan5480
Abstract
The faithful inheritance of the epigenome is critical for cells to maintain gene expression programs and cellular identity across cell divisions. We mapped strand-specific DNA methylation after replication forks and show maintenance of the vast majority of the DNA methylome within 20 minutes of replication and inheritance of some hemimethylated CpG dinucleotides (hemiCpGs). Mapping the nascent DNA methylome targeted by each of the three DNA methyltransferases (DNMTs) reveals interactions between DNMTs and substrate daughter cytosines en route to maintenance methylation or hemimethylation. Finally, we show the inheritance of hemiCpGs at short regions flanking CCCTC-binding factor (CTCF)/cohesin binding sites in pluripotent cells. Elimination of hemimethylation causes reduced frequency of chromatin interactions emanating from these sites, suggesting a role for hemimethylation as a stable epigenetic mark regulating CTCF-mediated chromatin interactions.
© 2018 Phys.org