A new mechanism involved in Staphylococcus aureus virulence and antibiotic resistance
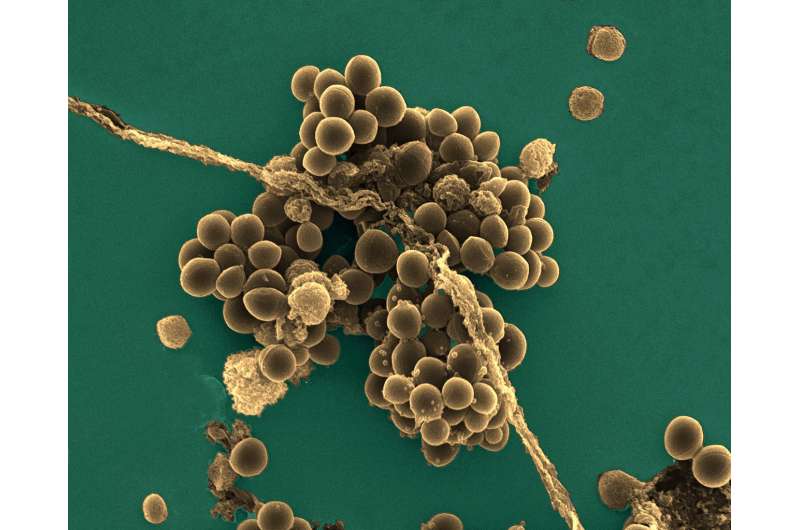
An Institut Pasteur-CNRS research team has characterized a Staphylococcus aureus gene involved in virulence, biofilm formation and resistance to certain antibiotics. These results open up new avenues for understanding the control of S. aureus virulence mechanisms. This work was recently published in the journal PLoS Pathogens.
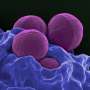
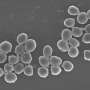
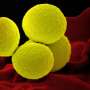
Staphylococcus aureus is part of the natural skin flora, preferentially colonizing external mucosa in 30 to 50 percent of the population, healthy carriers who develop no symptoms. But it is also a major human pathogen, causing diseases ranging from skin lesions (boils, impetigo, etc.) to endocarditis, acute pneumonia, osteomyelitis or sepsis. It is the leading Gram-positive bacterium responsible for hospital-acquired infections. The most dangerous strains are those that display resistance to multiple antibiotics. Such is the case of methicillin-resistant MRSA, widespread in hospitals and posing a major public health concern.
A team led by Tarek Msadek, a researcher at the Institut Pasteur (CNRS ERL 3526), is studying bacterial responses to environmental variations and their role in Staphylococcus aureus pathogenesis and host interactions. These responses are often genetically controlled by so-called "two-component" systems. During the study of one such system called WalKR, which is essential for bacterial survival, they characterized an additional component, SpdC, a membrane protein whose role was unknown. This component interacts with the WalKR system to control its activity, and its absence leads to a strong decrease in virulence, biofilm formation (bacterial aggregates), and resistance to certain antibiotics.
These results suggest that inhibition of SpdC could be used as an approach to combat S. aureus infections and understand the mechanisms involved in its transition from commensal to pathogen.
More information: Olivier Poupel et al, SpdC, a novel virulence factor, controls histidine kinase activity in Staphylococcus aureus, PLOS Pathogens (2018). DOI: 10.1371/journal.ppat.1006917