May 2, 2018 weblog
How to analyze your genome; Part I—Mitochondrial DNA
Genome analysis today is basically blind. It typically proceeds by randomly inspecting a smattering of possible variants that are only loosely associated with some disease or physical trait. Unless you already have a major health problem, this kind of narrowly focused crapshoot is not likely to be a game changer for you.
In this void, we would like to offer a more methodical approach—a simple formula to logically parse genomes at their critical points, and establish reliable physiologic predictors that are relevant for anyone. But first, a little background is required.
Our genomes are hybrids that have been built by viruses and bacteria. We have two of them, a big genome and a little one. Bioinformatics in general has mostly ignored the simple 16500 position mitochondrial DNA (mtDNA), and instead focused almost exclusively on the complex 3 billion position nuclear DNA (nucDNA). As we shall see, this is completely backward.
Except a small peptide called humanin, all the proteins that are coded within the tiny human mitogenome are used exclusively in respiratory complexes I through V. Two dueling 3-D print heads (see below) localized on either side of the double mitochondrial membrane system cooperatively extrude and deploy each subunit into the proper assembly compartment. Once there, assembly factors specific for each complex sequentially stitch each together. The mitochondrial matrix side of this membrane hosts the mitoribosomes while the cell side hosts the cytosolic ribosomes.
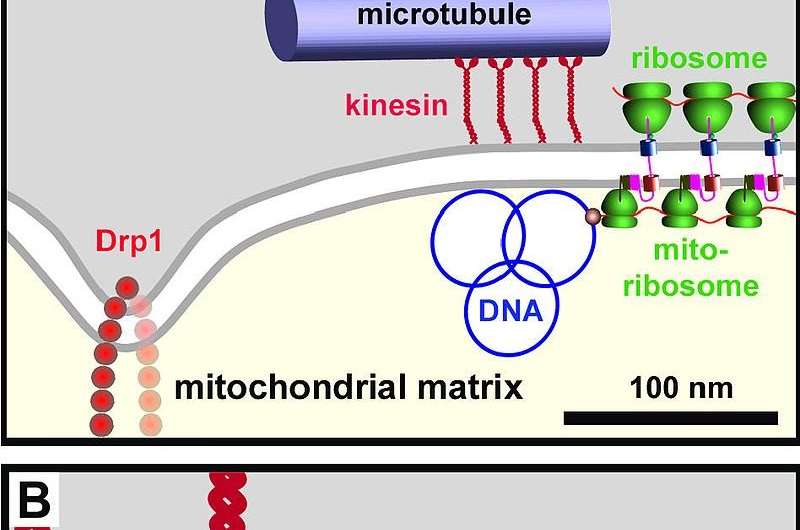
Mitochondria evolved as bacterial endosymbionts. They built the very first eukaryotic nucleus, and every nucleus since, using copy-paste-modify hardware they borrowed from viruses. Nucleotide tapes are read, written, and altered using mitochondrial-specific DNA and RNA polymerases that have long since been offloaded to the nucleus for storage, along with most of their other essential proteins. Understanding this mitochondrial construction project will be our key to unlocking the entire genome.
There are at least 1500 places in the nucDNA that we are concerned with. These are the locations that code for proteins, and some RNAs, that are used in mitochondria. The trick to recognizing these genes is that they usually start off with a mitochondrial localization sequence that targets them to the right organelle. Not all of these genes that have been culturally assimilated into the nucleus are migrants from the nucleoid. Many have simply duplicated themselves from existing nuclear genes and subsequently conjured up an alternative way to splice in an organelle localization motif.
While the full list of these genes (the "mitonuclear genome"), has yet to be completely discovered, many that have been mined so far reside online at the MitoCharta website. Researchers continue to find many more citizens of the mitonuclear genome. These are not expressed by all tissues, and do not always contain localization motifs, but can make their way in to mitochondria. Collectively, these are the critical points of the hybrid genome.
In other words, the sweet spot in genomics lies at the places where the two genomes intersect. To analyze them, we must search for correlations between polymorphism in the expanding mitonuclear genome and the tiny mitogenome. More specifically, we need to create a panel of effects that might be expected when variants in the >1500-strong mitonuclear genome are found alongside specific variants in the 13-strong mitogenome.
Until recently, searching disease databases for single variants that might match a patient's was the only way to analyze risk or diagnose many rare disorders. Mitochondrial disease, once considered extremely rare, is actually something that affects everyone in some form or another. While it is possible to create cell hybrids or 'cybrids' to explore effects of specific mtDNA mutations in a research setting, this is not typically done for individuals. Fortunately, there is now another way forward, namely, structural modeling and molecular dynamic simulation.
The beauty of this approach is that it can outperform crude database searches of other people's business (and frequently other organisms) that only return a diffuse hodgepodge of poor matches to an individual's particular set of variants. Now, any genetic profile can be directly reverse-engineered. In other words, we can individually tag all our variants, and then explore the implications of each on a complex-by-complex basis.
Each complex is embedded within a periphery of associated import, replication, translation, and metabolic cycle components of the mitonuclear arsenal that organizes into different macro-complexes under different conditions. In addition to mutations that alter core catalytic activities of individual subunits, it is now widely appreciated that changes in the peripheral amino acids where subunits interact often give the most readily visible effects. These border amino acids are the ones that most directly control assembly of subunits into higher-order structures—all the way up to the elusive respiratory supercomplex.
A good way to begin the analysis is to use an example of real mtDNA. I recently obtained a whole genome sequencing (WGS) from Dante Labs with a special request for mtDNA sequencing that cost only a few hundred dollars. I received the mtDNA results after a few weeks in the form of two files in a common format known as FASTQ. Each file contains the sequencing data as obtained by sequencing fragments of DNA in one direction. To compare the results with the Cambridge Reference Mitochondrial Sequence (CRS), and extract my particular variants, I uploaded my files at a website called mtDNA-Server , and used 'single ended' for file type.
In addition to listing your variants, the most abundant 'homoplasmic' sequence results from your submitted sample, you also receive results about heteroplasmy. This data consists of additional low-abundance reads, mainly from the so-called nuclear mitochondrial DNA segments (NUMTs). These are mostly nonfunctional relic copies of mtDNA residing on many chromosomes. Going forward, it will be important to analyze mtDNA from compartments other than saliva to get a better handle on actual heteroplasmy within different mitochondria.
For example, cardiac muscle cells or fibroblasts can preferentially accumulate mutated and deleted mtDNA. In other cases, different tissues deliberately maintain separate populations of heteroplasmic mitochondria. In my case, I received the following homoplasmic variants:
11788 C>T MT-ND4
1438 A>G MT-CYB (should be MT-RNR2)
15326 A>G MT-CYB
16519 T>C MT-DLOOP1
263 A>G MT-DLOOP2
4769 A>G MT-ND2
750 A>G MT-RNR1
8860 A>G MT-ATP6
The good news, here, is that while not everyone is a winner in the potluck of genetic recombination, everyone with mitochondria can be assigned to a particular haplotype. This tells them roughly where they fit in the human family tree, and to a lesser degree, the kinds of environments to which their mitochondria are adapted. Using tools like MitoMap and Phylotree, and the expert assistance of researchers Marie Lott and Shiping Zhang at the Children's Hospital of Pennsylvania, I found that I fall squarely within haplogroup "H." This was because I share all of the very common 263G, 750G, 1438G, 4769G, 8860G, 15326G, and 16519C markers with this group. The one rare allele I have, at 11788T, is only found in 25 of the 45,000 sequences in the MitoMap database and this bumps me up into the H56 subgroup. This haplotype is a predominantly Northern European subgroup believed to have originated shortly after the last Ice Age.
One thing that jumps out to me right away is an apparent abundance of A>G variants in my workup. A>G also seems to be overrepresented among known severe mitochondrial disease mutations, like, for example, MELAS. Marie and Shiping noted that while A>G is definitely a statistically favored transition ( G>A vs A>G shifts occur at a ratio of 2.26 to 1 ), their abundance can be partly explained by the prevalence of transitions over transversions and a natural strand asymmetry within mtDNA; The light strand, from which the mitogenome is numbered, has ~30 percent A, but only ~13 percent G. Furthermore, A and G variants are known to be preferentially localized to specific regions of the mtDNA.
Going down the variant list, we find that some protein-coding variants, like the 4769A>G in MT-ND2 or the 11788 C>T in the MT-ND4 gene were "synonymous." These genes code for subunits of NADH dehydrogenase complex I. Synonymous means that although the nucleotide changes, the new codon will still be recognized by the same mt-tRNA or family of similar tRNAs. Therefore, these variants are not as likely to affect the protein sequence itself. It is still possible that some minor changes to the speed or fidelity of translation may occur.
The 8860 A>G transition MT-APT6, on the other hand, is a nonsynonymous substitution. More specifically, this missense mutation from A >G changes the codon specificity from that of T to A. It is important to realize that in the codon world T is the amino acid threonine not the nucleoside thymidine, while A is alanine, not adenosine. To determine where this mutation occurs in the F-ATPase protein 6 subunit of complex V, we can use Mitomap to convert the 8860 mtDNA-referenced coordinate to protein-referenced coordinate of 112.
Using protein structure databases like Uniprot and interactive modeling sites like Protein Model Portal, we can punch in the alanine variant at position 112 and see where it lies among the various loops and folds of the protein. The main article image at top shows the whole ATP6 protein and the image below highlights position 112.
Searching the dbSNP Short Nucleotide Variations database revealed that several MT-CYB variants are associated with hypertrophic cardiomyopathy, so I wanted to do some more in-depth analysis there. One paper, which included my particular 15326A>G variant in their study, provides an excellent recipe. The authors of this 2013 study used PolyPhen to analyze mutations close to the conserved function heme-binding redox centers of MT-CYB. Although Polyphen offers feelings-based metrics that score various mutations as "possibly very damaging," it can also attribute these subjective descriptions to actual functional parameters like overpacking at buried sites in the protein.
Depending on whether or not crystal structures of human proteins are available at sufficiently high resolution, one can use prediction software like I-TASSER and Swiss-PDB Viewer to introduce amino acid changes and evaluate all possible rotamers. Changes in hydrogen bonding, macromolecular interactions and energy minimization can be performed using GROMOS force field.
Full molecular dynamics simulations are still not for the faint of heart. The NAMD and CHARMM22 programs, for example, generally require a lot of computing power. The MT-Cyb subunit interacts with several of the 11 subunits within complex III. Packages like the STRIDE simulator can calculate secondary structure at successive timepoints to reveal where interacting alpha helices transition and alternately transition to random coils to disrupt the structure of the complex.
We will wrap up our analysis of the mitogenome in the next article in this series, and also start to look at the mitonuclear genome as it becomes available to me. I am trying to follow in the footsteps of an early pioneer of open genomics, Brian Pardy, who was one of the first people to make his entire genome available for free to anyone interested in using it for analysis. My own information and files are available as they come in at this blog.
More information: us.dantelabs.com/
© 2018 Medical Xpress