Microbiome search engine can increase efficiency in disease detection and diagnosis
Big data makes big promises when it comes to providing insights into human behavior and health. The problem is how to harness the information it provides in an efficient manner. An international team of researchers has proposed a microbiome search-based method, via Microbiome Search Engine, to analyze the wealth of available health data to detect and diagnose human diseases.
They published their method on March 17, 2020 in mSystems, a peer-reviewed open access journal of the American Society for Microbiology. "Microbiome-based disease classification depends on well-validated, disease-specific models or markers," said Dr. Su Xiaoquan, from Single-Cell Center at Qingdao Institute of Bioenergy and Bioprocess Technology (QIBEBT) of the Chinese Academy of Sciences (CAS). "However, current models are lacking that information for many diseases."
In addition, SU said, multiple diseases can share the same biomarkers—the microorganisms that indicate something out of the ordinary, such as a mutated protein found in cancer cells, making it harder for researchers to correctly classify each one.
To combat these issues for disease detection and classification, Su and his joint software team from Single-Cell Center, QIBEBT and Center for Microbiome Innovation (CMI), University of California at San Diego (UCSD), developed a new search approach based on the whole microbial community a human body contains, collectively called the microbiome. Every person has a microbiome, even if they do not have a disease.
Traditional models compare samples from healthy subjects to those from people known to have specific diseases. With the new method, by searching based on specific outliers, rather than known biomarkers that can code for several diseases, the researchers can identify the microbiome state associated with the disease across different cohorts or sequencing platforms.
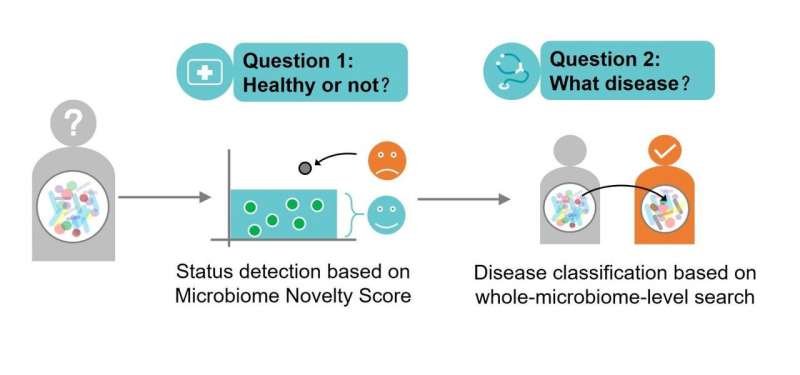
In this new approach, the research team employs a two-step process to identify disease. First, they search a baseline database of healthy individuals to detect any specific microbiome outlier novelty—or any known anomaly that differentiates the microbiome from a healthy state. They then search for that outlier in a database of disease-specific examples.
"Our strategy's precision, sensitivity and speed outperform model-based approaches," Su said.
The results of the search can provide quick predictions to help clinicians diagnose and treat diseases.
"This search-based strategy shows promise as an important first step in microbiome big data-based diagnosis," according to Rob Knight, Director of CMI and UCSD. "In light of the general shift of microbiome-sequencing focus from healthy to diseased hosts, the findings here advocate for adding more baseline samples from across different geographic locations."
Xu Jian, Director of Single-Cell Center, QIBEBT, agrees. Next, the joint Sino-U.S. team is working towards encouraging their colleagues to join a coordinated effort to continue expanding the microbiome database, to include every population and every ecosystem on the globe.
"With Microbiome Search Engine, performing a search can become as standard and enabling for new microbiome studies as performing a BLAST against your new DNA sequence is today." Xu said.
More information: Xiaoquan Su et al, Multiple-Disease Detection and Classification across Cohorts via Microbiome Search, mSystems (2020). DOI: 10.1128/mSystems.00150-20
Microbiome Search Engine: mse.ac.cn