Powerful sequencing technology sheds light on D.C.'s HIV epidemic
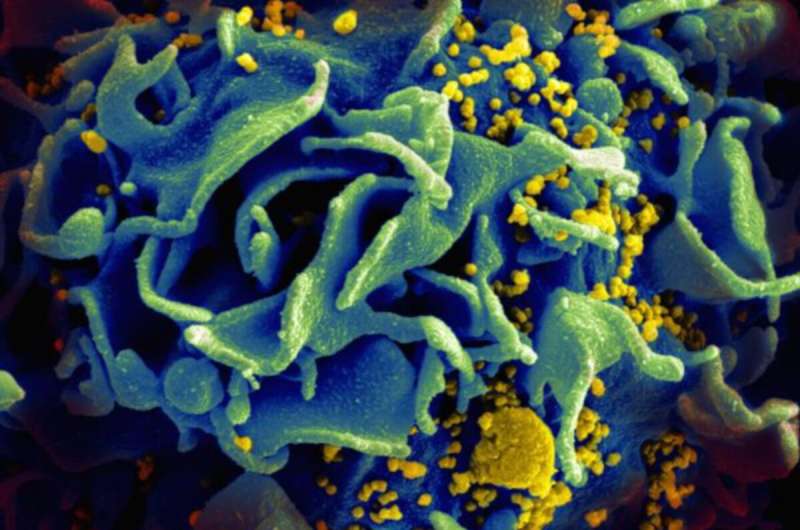
Despite significant progress against HIV/AIDS, the nation's capital is still battling an HIV epidemic with rates that are five times higher than the national average. A recent study by Milken Institute School of Public Health researchers at George Washington University uses powerful next-generation sequencing technology to learn more about how the virus is spreading and developing drug resistance in the District of Columbia.
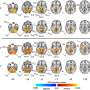
Researchers with GW's Computational Biology Institute (CBI) received blood samples from 68 people living with HIV in the District through collaborations with the DC Cohort. They sequenced the genetic material taken from the virus and found that HIV in D.C. is highly diverse—meaning there are many different strains in the local population. The authors theorize this diversity stems from D.C."s position as an international hub with a lot of temporary residents and visitors.
The researchers also found that about half of the study participants had at least one drug-resistant HIV strain. This information could be used to help provide targeted, more effective drugs to people with resistant HIV.
"Our findings of a genetically diverse and complex HIV epidemic in D.C. are scientifically important," said Keylie M. Gibson, who recently defended her dissertation at GW and lead author of the study. "At the same time, this study allowed us to give back to the District and the community through public health outreach and collaborations with organizations such as the DC Cohort."
Different subtypes of HIV are most prevalent in various parts of the world. Subtype B is most common in North America and Europe. Knowledge about which subtypes are common in a particular geographical location helps doctors identify the most effective treatment for individual patients.
"Ideally we want doctors to be aware that there might be different drug-resistant mutations circulating, and they might come across something they are not used to seeing or not very prevalent," Dr. Gibson said. "That knowledge could lead them to prescribe different drugs or at least be open to the idea that a patient might need a different drug combination based on drug resistant mutation profiles in different subtypes."
The study was carried out by a team of bioinformatics scientists and epidemiologists that included Dr. Gibson and Marcos Pérez-Losada, an assistant professor in the CBI. Using next generation sequencing technology along with epidemiological data, the team was able to map how different HIV strains were transmitted in the District. Learning more about the factors involved in such networks can be used to prevent new cases from occurring and to provide effective treatment to those already infected, the authors said.
"Our findings give us important clues about the spread, diversity and evolution of HIV in Washington, D.C.," said Dr. Pérez-Losada. "Such knowledge can be used to devise better methods for HIV prevention and treatment."
The study was published in the Feb. 6 issue of Scientific Reports, an online, open-access journal from the publishers of Nature.
More information: A cross-sectional study to characterize local HIV-1 dynamics in Washington, DC using next-generation sequencing, Scientific Reports (2020). DOI: 10.1038/s41598-020-58410-y