Researchers show how mutations in DNA packaging machines cause cancer
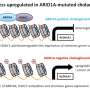
Like wrenches made of Legos, SWI/SNF chromatin remodeling complexes tighten or loosen DNA in our cells to control how genes are turned on and made into proteins. When assembled correctly, these complexes play a crucial role in the development of normal tissues, and when broken, they can lead to the development of cancer. These complexes are commonly disrupted by mutations in the genes that encode them—but how this leads to cancer is poorly understood.
New research from the Children's Medical Center Research Institute at UT Southwestern (CRI) determined how mutations in two key SWI/SNF proteins, ARID1A and ARID1B, can drive cancer development by disrupting the assembly of SWI/SNF complexes. The study, published in Nature Cancer, addresses fundamental questions about SWI/SNF biology as well as therapeutic strategies designed to kill cancer cells by targeting this complex.
"While it is abundantly clear that SWI/SNF components are defective in almost all cancer types, it is still fuzzy how mutations in components lead to broken SWI/SNF complexes, and how broken complexes cause disease," says study leader Hao Zhu, M.D., an associate professor at CRI. "In this study, we tried to cleanly break one important type of SWI/SNF complex to study how it falls apart, and how this leads to uncontrolled cancer growth."
SWI/SNF protein complexes help to pack and unpack DNA in the genome and are composed of 10-15 interacting proteins that can be arranged into different configurations in different tissues. Three main types of SWI/SNF complexes have been identified: cBAF, pBAF and ncBAF. But the roles they play in tissue development and disease have been unclear. To understand the importance of these complexes in animals, researchers at CRI focused on the cBAF complex. This complex was chosen because it is the most abundant one, and a subunit unique to this complex, ARID1A, is one of the most mutated genes in human cancer.
ARID1A is closely related to another protein known as ARID1B, which is also unique to cBAF. It has been shown that some cancer cells need at least one ARID1 protein to survive. To examine whether simultaneous loss of both ARID1A and ARID1B would be more likely to cause or kill cancer cells, researchers eliminated or knocked out both genes in mice. Strikingly, the loss of both ARID1A and ARID1B genes resulted in aggressive liver and skin cancer formation within weeks.
"In cancers where ARID1A is gone or mutated, one proposed strategy to stop cancer growth is to inhibit the replacement protein ARID1B. This method was predicted to kill cancer cells that might need cBAF function to survive," says Zhu. "However, our findings suggest that therapeutically targeting ARID1B could make matters worse by accelerating aggressive cancer development."
Researchers discovered that loss of these proteins led to the disassembly of the cBAF complex into many nonfunctional pieces.
They were able to uncover how ARID1A and ARID1B proteins maintain stabilizing connections between different components within cBAF complexes. This helped them pinpoint a number of important regions within these ARID1 proteins, that when mutated can make cBAF complexes fall apart. Interestingly, the importance of these regions also explains why mutations accumulate in these regions in human cancers. When cBAF falls apart, the leftover components interfere with the composition and function of other types of SWI/SNF complexes, which further contributes to cancer.
"We hope that the findings in our paper will change the way people think about the molecular consequences of SWI/SNF disruption and how mutations in this complex drive malignancy," says Zixi Wang, Ph.D., a postdoctoral researcher at CRI, assistant instructor of pediatrics at UTSW, and lead author of the paper.
More information: Dual ARID1A/ARID1B loss leads to rapid carcinogenesis and disruptive redistribution of BAF complexes, Nature Cancer (2020). DOI: 10.1038/s43018-020-00109-0 , www.nature.com/articles/s43018-020-00109-0