New AI-based image processing algorithm for accurate cervical cancer detection
Cervical cancer affects more than half a million women annually causing more than 300,000 deaths. Detection of cancer in its early stages is essential to eradicate the disease from the patient's body. However, regular population-wise screening of cancer is limited by its expensive and labor-intensive detection process. To detect malignancy, clinicians need to classify individual cells from a stained slide, consisting of more than 100,000 cervical cells.
Thus, Computer-Aided Diagnosis (CAD) systems are a viable alternative for easy and fast cancer detection.
"Traditional machine learning-based methods, although computationally less complex, require extraction of handcrafted features, and feature selection for classification," says Dmitrii Kaplun, associate professor at the Department of Automation and Control Processes, Saint Petersburg Electrotechnical University. "This limits the performance of such models because of the two main reasons: extraction of handcrafted features becomes difficult for complex data pattern, and all these features may not be sufficiently informative, thus adversely affecting the model's performance."
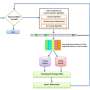
Keeping this in mind, the researchers proposed a new technique. They formed an ensemble-based classification model, using three Convolutional Neural Network (CNN) architectures—Inception v3, Xception and DenseNet-169. Ensemble learning is a strategy that entails considering more than two models to make a final prediction. Neural networks are pre-trained on ImageNet dataset for Pap stained single cell and whole-slide image classification.
"The proposed ensemble scheme uses a fuzzy rank-based fusion of classifiers by considering two non-linear functions on the decision scores generated by said base learners," says Kaplun. "Unlike the simple fusion schemes that exist in the literature, the proposed ensemble technique makes the final predictions on the test samples by taking into consideration the confidence in the predictions of the base classifiers."
The proposed model has been evaluated on two publicly available benchmark datasets, namely, SIPaKMeD Pap Smear and Mendeley Liquid Based Cytology (LBC) dataset, using a 5-fold cross-validation scheme. On the SIPaKMeD Pap Smear dataset, the proposed framework achieves a classification accuracy of 98.55% and sensitivity of 98.52% in its 2-class setting, and 95.43% accuracy and 98.52% sensitivity in its 5-class setting. On the Mendeley LBC dataset, the accuracy achieved is 99.23% and sensitivity of 99.23%.
The results were published in the journal Scientific Reports. The proposed model seems to outperform many of the state-of-the-art ones, justifying its effectiveness.
This quick-identification tool can be used as a plug-and-play model. It requires minimal support from clinicians to perform a cervical cancer screening. Therefore, it is suitable for use in this field of medicine.
The researchers are expecting to try contrast enhancement techniques or prior segmentation of cells for more accurate classification in cases of poor image contrast or in the presence of overlapping cells. They also may consider ensembles of other base learners, and explore different rank generation functions to perform the ensemble.
More information: Ankur Manna et al, A fuzzy rank-based ensemble of CNN models for classification of cervical cytology, Scientific Reports (2021). DOI: 10.1038/s41598-021-93783-8