Incidence estimation from SARS-CoV-2 genomes
A team of researchers from the Robert Koch Institute in Berlin and the Max Planck Institute for the Science of Human History present a novel method for the rapid estimation of incidence trajectories from serially sampled SARS-CoV-2 genomes and validate it using phylodynamic methods.
Daily counts of new COVID-19 cases remain a basis for evaluating the state of the pandemic and are vital for making informed decisions on public interventions. These case counts, however, are based on positive diagnostic test results and are thus highly dependent on the underlying testing strategy. But testing strategies differ from region to region and have drastically changed over time, making their precise influence on the number of daily new diagnoses hard to predict. More robust estimates of the number of newly infected individuals are therefore necessary for pandemic surveillance.
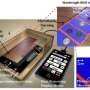
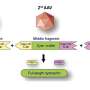
In order to better estimate new infection rates, researchers have developed and tested a new computational method that infers temporal profiles of viral incidence purely from genomic sequences and their sampling dates. Because the viral genome underlies a steady mutation process, the changes in its sequence through time also track its spread through the population.
"The mutations that arise in the viral genomes leave a signal that allows us to connect genetic diversity with viral population size and, in this case, also incidence," says Denise Kühnert, co-author of the study.
Effect of non-pharmaceutical interventions
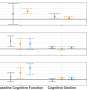
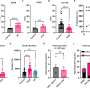
By calculating piece-wise constant estimates of the effective reproductive number between significant changes in non-pharmaceutical interventions, the study highlights potential effects of public measures on the spread of COVID-19.
The lockdown measures taken by many European countries are a telling example. In the Spring of 2020, after the implementation of strict lockdown measures in Europe, the effective reproductive number fell sharply to below 1. When these measures were lifted at the beginning of Summer 2020 in most countries, the effective reproductive number increased to greater than 1.
Phases of case under-detection
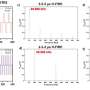
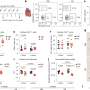
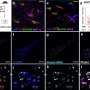
The team conducted extensive state-of-the-art phylodynamic analyses using SARS-CoV-2 genomes from four different regions (Denmark, Scotland, Switzerland and the Australian state Victoria) to validate the new method.
"By comparing our results with reported case numbers, we were able to detect periods of underreporting," says Ariane Weber, Ph.D. student in the research group and co-author of the study. "These include the first wave of infections in Scotland and Victoria, as well as small epidemic waves in Europe in the summer of 2020, which are not visible in the diagnostic case counts," Weber adds.
Relative case detection
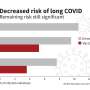
By using the incidence correlate generated by the new method and public data on deployed testing strategies, the researchers were also able to calculate case detection rates through time. These suggest that an increase in testing capacity generally leads to higher proportions of detected cases. Surprisingly, however, the detection probability decreases when the criteria for testing are relaxed. This provides a potential explanation for the observed under-detection of cases in Europe in Summer 2020, when looser testing criteria were combined with unchanged test capacities.
The study is published in Nature Communications.
More information: Rapid incidence estimation from SARS-CoV-2 genomes reveals decreased case detection in Europe during summer 2020, Nature Communications, DOI: 10.1038/s41467-021-26267-y