New imaging method helps to provide new, more personalized cancer treatments

As cancer therapy has advanced, the complexity of oncology has become apparent. There are many moving parts in a cell that can malfunction leading to a cancerous path, one example being epidermal growth factor receptors (EGFR)—a group of proteins that control cell growth. It includes HER1 (EGFR), HER2, HER3 and HER4.
If EGFRs are mutated, they can lead to the development of numerous cancers. A common problem with EGFRs is dimerization—when two receptors near each other (e.g. EGFR and HER3, HER2 and HER3) interact to accelerate tumor growth. Dimerization can positively and negatively impact the ability of EGFR treatments to act against the tumor, depending on certain factors such as the treatment being used or the type of tumor. Due to this role, dimerization is a good measure for cancer therapy effectiveness. Analyzing these interactions are problematic, however, as they're too small to detect with conventional light microscopes.
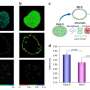
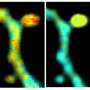
To combat this, King's researchers have worked closely with a new imaging method called FLIM-FRET that combines two methods. The FLIM method measures the rate of decay from fluorophores (a molecule that re-emits any light absorbed) added in the FRET method. One receptor is engineered with a donor fluorophore, the other receptor is given an acceptor fluorophore. If they're in proximity, the donor transmits energy to the acceptor fluorophore, making the acceptor fluoresce in a different color.
As FLIM measures the fluorophore's rate of decay, you can calculate the transfer rate between the two receptors. Combined, FLIM-FRET provides the tools to visualize and quantify dimerization interactions.
Led by Professors Tony Ng and Simon Ameer-Beg, researchers at King's have been using FLIM-FRET microscopes since 2001 to analyze protein-to-protein interactions in tumors, including EGFRs. King's researchers have been making their own developments them to improve its speed and accuracy, culminating in completing culminated the SWARM microscope in 2019.
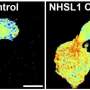
The new microscope has 1,024 beams for exciting and detecting fluorophores, making the process significantly quicker and improving resolution while also reducing errors. Research into assays has also allowed scientists to improve identification of target receptors and attach fluorophores. Because of these advances, King's researchers were able to analyze EGFR dimerization.
This newfound route of experimentation could change the lives of 20% of patients who receive a diagnosis for breast, HNSCC (head and neck squamous cell carcinoma), lung and colorectal cancers. Over 31,000 people per year could one day benefit from the insights that FLIM-FRET can offer. Many researchers, as well as new spinouts and pharmaceutical companies, are already collaborating with King's to use FLIM-FRET.
Daiichi Sankyo have been working with King's for seven years on several studies, such as a phase 2 trial studying the combination therapy Patritumab-Cetuximab in HNSCC. What unites their studies is using FLIM-FRET to better understand or/and target dimerization. Other companies, such as Roche, Incyte, UCB and AstraZeneca have also worked with King's to support or take advantage of FLIM-FRET.
It's clear that the innovations in FLIM-FRET developed by King's researchers have been a key part of our collective and continued fight against cancer. FLIM-FRET promises to be another tool that can help continue to fulfill the promise of personalized medicine and develop more therapeutic options against a range of cancers.