Developing AI tools for breast cancer precision pathology

Breast cancer is the most common cancer type in women globally, but there is a continuous decline in mortality rates. A partial explanation for this may be that effective treatments are being developed for subgroups of breast cancer patients. However, it is still challenging to optimize treatment plans for every individual.
In her thesis, Yinxi Wang at the Department of Medical Epidemiology and Biostatistics, aimed to improve disease outcome and to decrease the burden associated with unnecessary treatment and adverse drug effects. Yinxi aimed to develop artificial intelligence-based tools to improve individualized medicine for breast cancer patients.
What are the most important results in your thesis?
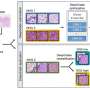
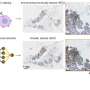
My thesis includes four studies where we developed deep learning models to improve histological grading and to build a scalable approach in predicting tumor average or spatial gene expression levels. We also sought to stratify breast cancer patients based on either histological patterns or intra-tumor gene expression heterogeneity for future personalized treatment regimens. We did take a few steps along the road, and our models that were optimized using whole slide images that are routinely collected for diagnostic purposes, making additional tissue sampling unnecessary, i.e., a cost-efficient solution compared to molecular diagnosis.
Why did you choose to study this particular area?
The field I'm working in is an interdisciplinary area called computational pathology, where we develop AI-based computational tools to facilitate cancer diagnosis and patient characterization.
The area attracts me mainly for two reasons: firstly, pathology is a very important domain that has long been the central part in diagnosing cancer; in addition, the morphological patterns that can be observed from the diagnostic tissue sections is a reflection of underneath intrinsic differences between individual tumors, thus, there is great volumes of information that can be extracted for improved precision medicine, and this possibility has made the area extremely fascinating to me. Secondly, the advances in technology, including the novel development of AI-based modeling has enabled rapid digitization and analysis of large numbers of tissue slides, making it possible to extract crucial information by integrating substantial amounts of pathological data and to further facilitate precision pathology by building powerful quantitative solutions. Performing statistical and machine learning modeling for this aim has been the most intriguing research field for me.
What do you think should be done moving forward in this research area?
On the one hand, deep learning remains a powerful tool that has the potential to help solving many clinically remaining questions; on the other hand, more studies have to be carried out to strengthen the performance of such models in real-world applications.
For the first aspect, there is lots of exciting research that can be carried out. For instance, it is now possible to combine information including radiological and histological image data as well as molecular data to obtain a more comprehensive understanding towards individual tumor, thus further facilitating precision medicine.
For the second aspect, we still need to find ways to improve the model's generalizability when handling domain shift or abnormal instances; we also need large scale validation studies to gather more evidence regarding the potential benefit of proposed methodologies.
More information: Artificial intelligence for breast cancer precision pathology. openarchive.ki.se/xmlui/handle/10616/48327