This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Study identifies novel genetic causes of male infertility
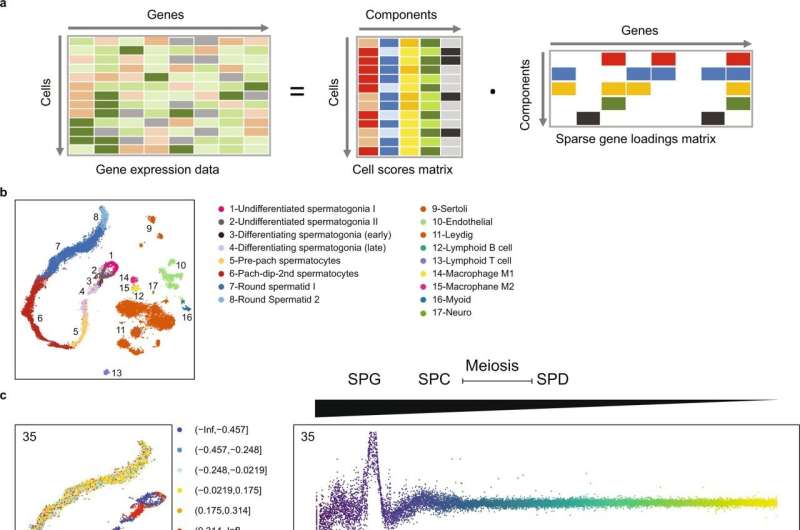
A recent study published in Nature Communications has identified novel genetic causes of non-obstructive azoospermia (NOA), the most severe form of male infertility, findings that improve the characterization of the disorder and may inform future treatment strategies and interventions.
NOA occurs in 1% of all men and is virtually incurable. Except in cases where a person has undergone radiation or chemotherapy treatment, the underlying etiology of NOA usually remains unknown.
"This is important because if a man was known to have an underlying genetic etiology of NOA, you would know to not do a sperm extraction procedure to try to retrieve sperm for fertility purposes," said Emily Jungheim, MD, the Edmond Confino, MD, Professor of Obstetrics and Gynecology, chief of Reproductive Endocrinology and Infertility in the Department of Obstetrics and Gynecology and a co-author of the study.
In the current study, investigators performed sequenced exomes—the coding portions of genes—in DNA samples from more than 1,000 men diagnosed with NOA who were enrolled in the GEnetics of Male INfertility Initiative (GEMINI) study.
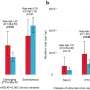
In 20% of these men, the investigators identified more than 200 genes with known roles in sperm production and after integrating single-cell RNA sequencing data, they also found more than 20 molecular subgroups of NOA genes with synchronized gene expression patterns. In mouse models of NOA, they also similar molecular subgroups of these genes, confirming their findings.
Ultimately, understanding the underlying genetic etiologies of reproductive diseases in both men and women can help clinicians know when and how to apply treatments more efficiently, more effectively and at lower costs, according to Jungheim.
"Genetic testing in reproductive medicine is largely limited to testing embryos—we have very little diagnostic testing available for people who fall short of making embryos. We are often left throwing our hands up and trying treatment after treatment until we have nothing left to offer except donor gametes. It would be better for everyone if we could provide answers instead," Jungheim said.
More information: Liina Nagirnaja et al, Diverse monogenic subforms of human spermatogenic failure, Nature Communications (2022). DOI: 10.1038/s41467-022-35661-z