This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
Researchers test DNA editing, recommend steps to improve accuracy
By studying the effects of DNA variants between people, researchers can gain insights into the genetics of diseases. It allows them to tailor medical treatment to a person's genetic makeup.
Modeling variants in an experimental setup is a common way for researchers to gain this knowledge. The CRISPR-Cas9 system, a broadly used gene editing tool, allows researchers to precisely change the sequence of a genome and study the effects of that genetic variation on cellular function. Understanding genome editing experiments and improving the accuracy of their results is crucial to successful research.
In a recent study published in The CRISPR Journal, researchers from Mayo Clinic, Yale University, Oklahoma Medical Research University and Baylor College of Medicine investigated the CRISPR-Cas9 system.
A typical genome editing experiment consists of three steps:
- Editing cultured cells.
- Cloning cells and selection of clones with or without the intended edit.
- Comparison of clones with or without the intended edit.
A clonal line is a group of cells or organisms with the same genetic makeup. Clonal lines are essential in research as they provide a consistent and reproducible source of genetically identical cells or organisms for experimentation.
Clones with and without the intended edits are identical or closely similar genotypes except for the induced mutation.
"This approach is often highlighted as particularly rigorous and powerful in that it allows comparing the effect of single mutations in an otherwise identical genetic background," explains lead author Alexej Abyzov, Ph.D., a researcher in the Department of Quantitative Health Sciences at the Mayo Clinic Center for Individualized Medicine. "However, the genomes of selected clones can be different, violating the assumption of matching (except for an edit) genome in compared clones."
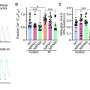
The team conducted three independent CRISPR-Cas9 editing experiments and used whole genome sequencing to analyze the extent of genomic differences in the clones. After expanding multiple clones, they examined somatic copy number alterations (i.e., large structural changes in the genome) and single nucleotide variants (i.e., point mutations) in each clone.
They rarely found off-target edits, a type of unintended genetic change occurring when gene editing tools mistakenly edit a gene or sequence that was not the intended target.
However, they detected hundreds to thousands of single nucleotide mutations distinctive to each clone. Clones also differed in copy number variations, such as deletions and duplications. Some of those were several kilobases (1,000 consecutive nucleotides) to megabases (1,000,000 consecutive nucleotides) in size.
"Copy number alterations represent the largest source of genomic divergence among clones," says Arijit Panda, Ph.D., a Mayo Clinic research fellow and first author of the study. "Our study shows that culture clones selected after DNA editing experiments do not have the same or closely similar genotypes and the gene's physical expressions may vary from the intended edits."
In order for researchers to correctly interpret DNA editing experiments and show a meaningful comparison between edited lines, the study's authors recommend:
- Screening clones for mutations and large copy number alterations acquired in culture to determine the largest source of genomic divergence between clonal lines.
- Excluding from consideration clones with large copy number alterations or mutations having potentially functional consequences.
- Comparing a mix of multiple unedited clones with that of edited clones to dilute the possible effect of mutations in each clone.
Dr. Abyzov says further studies are needed to determine the best and most cost-effective screening strategy before integrating these recommendations into CRISPR-Cas9 gene editing research at Mayo Clinic.
More information: Arijit Panda et al, Clonally Selected Lines After CRISPR-Cas Editing Are Not Isogenic, The CRISPR Journal (2023). DOI: 10.1089/crispr.2022.0050