This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Genomic sequencing method may help curtail syphilis spread
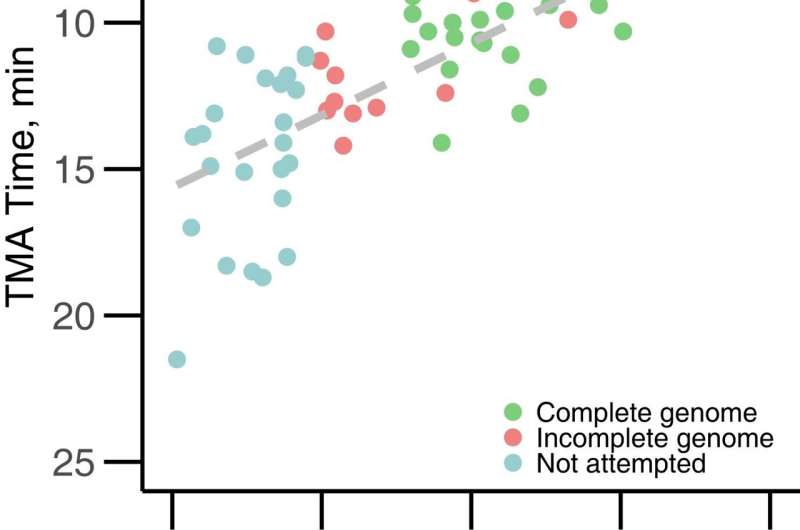
An easily implemented genome-sequencing method could help public health officials combat America's ongoing syphilis epidemic, researchers at the University of Washington School of Medicine and Public Health-Seattle & King County demonstrate in a new paper.
Syphilis rates in the United States are surging. Rates jumped nearly 32% in 2020-21 alone, while rates of congenital syphilis, which causes stillbirth and infant death, rose more than 219% from 2017 to 2021.
Genome sequencing determines the order of the DNA molecules that make up an organism's genetic code. Investigators can use this sequence as a genetic fingerprint to identify different strains and establish their relationships to other strains that might be circulating in a community.
Genome sequencing is commonly used to track other sexually transmitted diseases such as gonorrhea and chlamydia, but most public health labs do not use it to track syphilis.
One reason is that specimens of the bacterium that causes syphilis, Treponema pallidum, can be hard to obtain. Another is that the bacterium's genome changes relatively slowly. This makes it less likely that strains in a community will vary enough to merit such tracking during a fast-moving outbreak.
In the new study, however, the UW Medicine researchers and their local public health collaborators showed it is possible to sequence the bacterium's genome with standard equipment and to use the information to understand how different strains are spreading.
"Our workflow uses the same specimens and platforms used to detect gonorrhea and chlamydia, so it's very scalable and could be implemented by public health labs relatively quickly," said virologist Dr. Alex Greninger, an associate professor of laboratory medicine and pathology, who led the project.
The study findings are published online today, Sept. 28, in the Journal of Infectious Diseases. Nicole Lieberman, a senior research scientist in the Greninger lab, was the paper's first author. Here is a copy of the paper.
To test their approach, the UW researchers sequenced T. pallidum DNA found on swabs used to test 24 individuals for sexually transmitted infections. All the patients were seen at the Seattle Sexual Health Clinic.
The researchers determined that two clinically indistinguishable strains of syphilis were circulating in Seattle. Twelve of the patients were infected by strains belonging to the "Nichols lineage" and the other 12 by "SS14" lineage strains.
"When we looked at the demographics, we found clustering by sexual networks," Leiberman said. "The Nichols-lineage strains were predominantly found among men who have sex with men, while several heterosexual women and men were infected with a single SS14-lineage strains. This shows how this toolkit can be used to help public health authorities better understand how syphilis spreads within and between communities."
The genomic analysis also revealed that all the strains had acquired a mutation that made them resistant to the antibiotic azithromycin. Although most strains of T. pallidum remain treatable with penicillin, the finding that azithromycin-resistant strains are widespread in the community is concerning, Greninger said.
"That 100% of strains are resistant to this antibiotic, which isn't even prescribed by the Seattle Sexual Health Clinic, concentrates the mind," he said.
To show it is possible to use sequencing to detect subtle changes between strains that might arise during an outbreak, the researchers performed detailed deep sequencing of a gene called tprK. This gene codes for a protein found on the surface of the bacterium. The protein frequently changes to evade the body's immune response.
The researchers found that tprK sequences tended to remain closely matched in samples taken from the same patient and among samples from one cluster of heterosexual patients. Those findings suggest tprK could help identify closely related strains. This capability would be particularly useful in tracking outbreaks ongoing in communities.
"As demonstrated by the recent Listeria outbreak solved through sequencing in Washington state, pathogen genomics is increasingly routine in public health," Greninger said, "but there are still bugs like T. pallidum that are not well-covered by existing workflows and that is where UW Medicine can help push the field forward."
More information: Alex Greninger et al, Genomic epidemiology of Treponema pallidum and circulation of strains with diminished tprK antigen variation capability in Seattle, 2021-2022, The Journal of Infectious Diseases (2023). DOI: 10.1093/infdis/jiad368