This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
proofread
Precision treatment for pneumonia care: Metagenomic sequencing takes the lead
Community-acquired pneumonia (CAP) is a major infectious disease worldwide and contributes to high mortality and massive economic burden. Hospital mortality among the severe CAP (SCAP) remains high, ranging from 25% to more than 50%.
Early identification of patients at high risk of death is essential for improving patient outcomes. However, predicting outcomes in patients with SCAP is challenging, as the disease is complex and influenced by various factors, including the types of pathogen causing the infection, the host immune response, and underlying medical conditions.
In this study published in eBioMedicine, a team led by BGI Genomics Chief Science Officer of Infection Department Dr. Jinmin Ma and team led by Ruijin Hospital Prof. Jieming Qu conducted research on SCAP patients to explore the pulmonary microbiota and host responses of different outcomes since the genetic differences could be detected more easily in the most severe patients as opposed to mildly severe ones.
Methods and findings
BGI Genomics researchers used metagenomic and transcriptomic analysis to identify a new set of biomarkers that can predict 30-day mortality in patients with SCAP.
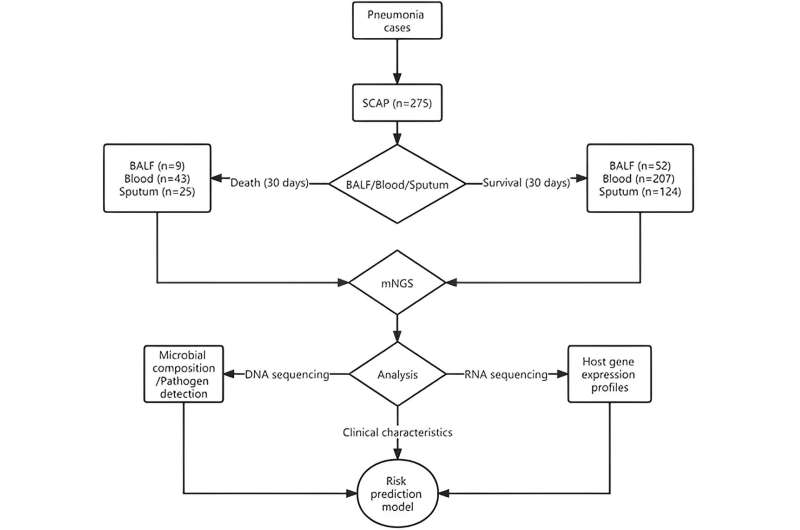
The study included 275 patients with SCAP from 18 hospitals in China.
Researchers performed DNA and RNA-based metagenomic next-generation sequencing of bronchoalveolar lavage fluid (BALF), sputum, and blood samples from 275 SCAP patients with varied characteristics and outcomes to analyze the differences in the microbes and host responses between them.
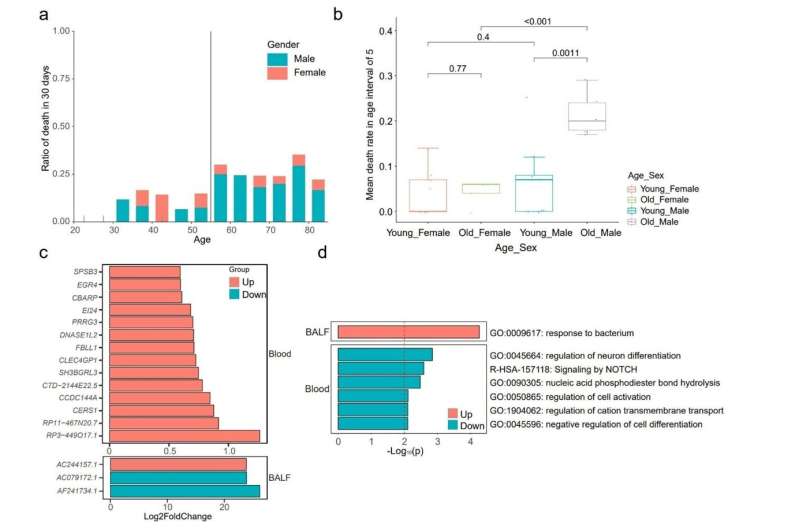
The researchers identified nine sets of biomarkers, both metagenomic and transcriptomic, that were associated with 30-day mortality.
The biomarkers were validated in an independent cohort of patients with SCAP and were able to predict 30-day mortality with an accuracy of 85%. This is significantly higher than the accuracy of existing clinical prediction models, which typically have accuracies of around 70%.
Other key findings:
- The study revealed that 30-day mortality was independent of pathogen category, microbial diversity or specific microbial taxa, while significant differences in host gene expression patterns were suggested to be responsible for different outcomes.
- Clinical characteristics analysis showed that male sex with age over 55 years was a risk factor for poor prognosis, and specific enrichment of genes and signaling pathways were found in omics data.
Potential of biomarkers utilization
More information: Jingya Zhao et al, A multicenter prospective study of comprehensive metagenomic and transcriptomic signatures for predicting outcomes of patients with severe community-acquired pneumonia, eBioMedicine (2023). DOI: 10.1016/j.ebiom.2023.104790