This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Sister cells uncover pre-existing resistant states in cancer
In many cancers, such as ovarian cancer, each round of chemotherapy kills the majority of cancer cells, while a small population of them survives through treatment. These cells are typically more resistant for the next cycle of therapy and can thus regrow to a deadly, treatment resistant tumor.
In a recent study published in Nature Communications, researchers at the University of Helsinki wanted to know how this small population of surviving cells differs from the other more sensitive cells already before the treatment. To enable this cellular time travel, they developed ReSisTrace, a methodology that takes advantage of the similarity of sister cells to trace back pre-existing treatment resistance in cancer.
Labeling cancer cells with genetic barcodes
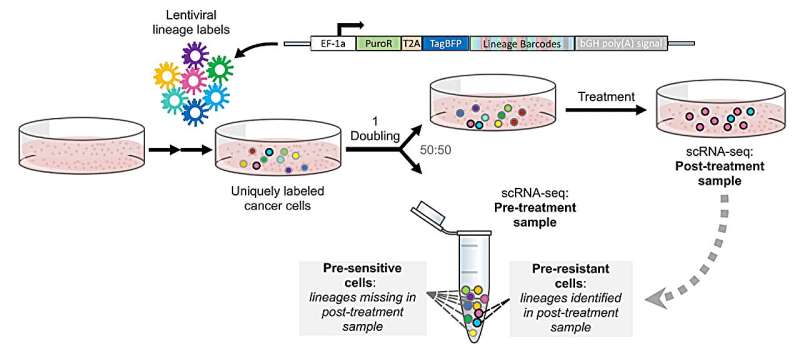
"In ReSisTrace, we label cancer cells uniquely with genetic barcodes and allow them to divide once, so that we get two identical sister cells that share the same barcode. We then analyze single-cell gene expression from half of the cells before the treatment, while treating the other half with chemotherapy, or other anti-cancer treatment. From the surviving cells we can identify the barcodes of resistant cells.
"Using their sister cells analyzed before the treatment, we can discover how the cells that will survive through treatment differ from the pre-sensitive cells, thus revealing the pre-existing resistant states," says Jun Dai, Ph.D. student in Anna Vähärautio's group, who developed the methodology to trace sister cells.
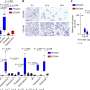
The method was applied to reveal resistant cell states against chemotherapy, targeted therapy or innate immunity in high-grade serous ovarian cancer.
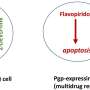
"We found that genes associated with proteostasis and mRNA surveillance are important to explain pre-existing treatment resistance. Interestingly, we found that DNA repair deficiency that is very common in ovarian cancer, sensitized these cells to not only chemotherapy and PARP inhibitors but also to NK killing," says Shuyu Zheng, a Ph.D. student from Jing Tang's group, who spearheaded the computational analysis.
Associate Professor Jing Tang's laboratory then leveraged the revealed gene expression changes to predict small molecules that could shift the cells from a resistant state to a sensitive state.
"We developed a computational method to correlate the resistant states with the gene expression changes induced by a drug. Ideally, if a drug can reverse the resistant cells' gene expression profiles, then it can be considered as a potential hit to overcome the resistance," says Associate Professor Jing Tang, and a team leader in Systems Oncology Research program, University of Helsinki.
Researchers found that most of the predicted small molecules indeed changed the gene expression patterns of cancer cells toward sensitive states. Most importantly, after adding these drugs, cancer cells were significantly more sensitive to carboplatin, PARP inhibitor or NK killing, illustrating that the pre-resistance states identified by ReSisTrace were functionally relevant and targetable.
"Our novel experimental-computational approach really leverages the power of single-cell omics and pharmacological data integration," Associate Professor Jing Tang says.
"The method we developed reveals the features of cells that will—in the future—become resistant to anti-cancer treatments by coupling cell state and fate in sister cell resolution. It is widely applicable to identify and target pre-existing resistant cell states across cancer types, as well as against different treatment modalities, including immunotherapies.
"Our approach paves the way for development of sequential cancer therapies that can block resistance before it even emerges," concludes Anna Vähärautio, K. Albin Johansson Cancer Research Fellow, Foundation for the Finnish Cancer Institute and a team leader in Systems Oncology Research program, University of Helsinki.
More information: Jun Dai et al, Tracing back primed resistance in cancer via sister cells, Nature Communications (2024). DOI: 10.1038/s41467-024-45478-7