This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
proofread
Researchers identify abrupt epigenetic aging of the colon
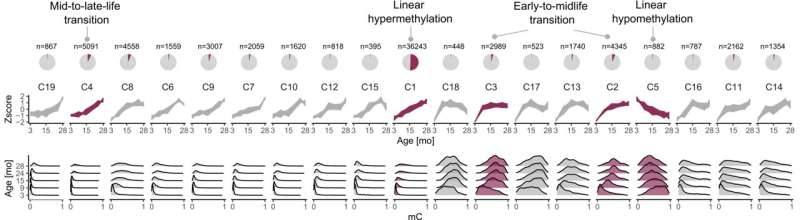
DNA methylation data provide extremely accurate age predictors, but so far, little is known about the dynamics of this epigenomic biomarker over the course of life.
A team of researchers from the Leibniz Institute on Aging–Fritz Lipmann Institute (FLI), the Jena University Hospital (UKJ), and Christian-Albrecht University of Kiel (CAU) led by Dr. Maja Olecka, Dr. Alena van Bömmel, Prof. Christoph Kaleta, Dr. Christiane Frahm, and Prof. Steve Hoffmann has now made a discovery significantly contributing to narrowing this knowledge gap. The bioinformaticians investigated the epigenetic patterns (methylation trajectories) of the male mouse colon at different stages of aging.
In addition to the continuous DNA methylation changes observed during aging, they noted significant hypermethylation events at distinct life stages. According to the study, published in Nature Communications, two key waves of epigenetic modifications take place; during the shift from early to middle age (3–9 months) and middle to late age (15–24 months). The life of rodents can thus be divided into three epigenetic stages.
The findings provide a new perspective on the dynamics of aging. "At least in the colon, the aging can no longer be regarded solely as a continuous accumulation of changes occurring during life. The identification of sudden methylation changes suggests that switch-like mechanisms also contribute to aging," explains Dr. van Bömmel, one of the two lead authors.
Her colleague Dr. Olecka adds, "Importantly, nonlinear methylation changes are likely to impact the organ function. Genes affected by sudden changes at both DNA methylation and gene expression level are important players in various aspects of colon health and function like intestinal barrier, colon cancer or enteric nervous system."
The research team also developed a clock-like classifier called STageR (STage of aging estimatoR), which can be used to predict the epigenetic stage of life in mice accurately. STageR differs from existing epigenetic clocks in that it is based on nonlinearly modified DNA regions and is not restricted to a fixed set of predictors. STageR thus expands the toolkit of machine learning models used in aging research. The accuracy of STageR has been confirmed in a separate strain of mice and on publicly available data.
The next steps of this exciting research project are already in sight. "The discovery of nonlinear methylation trajectories is a thrilling starting point for a new line of research. For example, we want to check whether the same phenomenon takes place in other tissues," explains Prof. Hoffmann.
More information: Maja Olecka et al, Nonlinear DNA methylation trajectories in aging male mice, Nature Communications (2024). DOI: 10.1038/s41467-024-47316-2