This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
A milestone in epigenetics of reproduction: Researchers crack the male fertility code
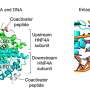
Each cell contains a set of instructions in its DNA that determines which genes are expressed and which are silenced. The correct programming of these so-called epigenetic programs, including DNA methylation, is essential for fertilization and development.
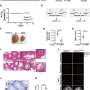
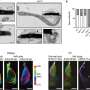
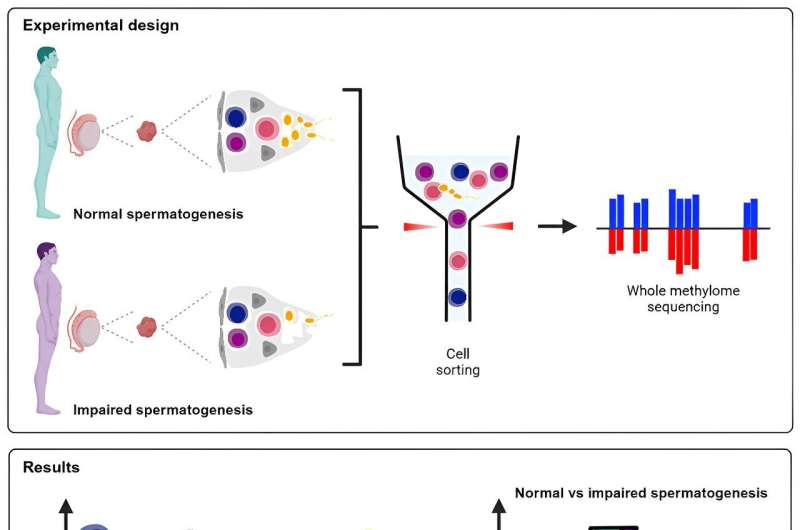
A team of researchers from the University of Münster have for the first time identified the DNA methylation program behind sperm production (spermatogenesis) in the human. They found that during spermatogenesis the entire genome is reprogrammed. Moreover, when they analyzed cells from infertile men, they found that some regions in the genome are not correctly programmed, revealing a new potential cause of male infertility.
The code has been cracked, at least the one leading to the production of sperm in humans. For this to be successful, the genes involved in the process need an "instruction manual." In other words, for the germline to generate sperm through the process of spermatogenesis, certain chemical patterns must be established in the DNA.
A research team led by Dr. Sandra Laurentino and Professor Nina Neuhaus from the Center of Reproductive Medicine and Andrology (CeRA) of the University of Münster Medical Faculty has now discovered the specific instructions behind this. On top of that, the Münster researchers also identified a new possible cause of male infertility based on faulty genome regulation. These results have been published in the American Journal of Human Genetics.
The translational study, led by biochemist Laurentino and biologist Neuhaus, focused on DNA methylation, a type of chemical modification in the DNA that regulates genes. This forms a sort of computer program in which genes of different cells are "switched on and off" in order for spermatogenesis to progress.
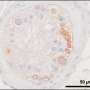
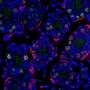
The testicle, where sperm is produced, is a very complex tissue, explains Dr. Laurentino. This is why the "instructions" behind spermatogenesis remained unknown until now.
The research group achieved a breakthrough with colleagues from the Max Planck Institute for Molecular Biomedicine in Münster, who are now working at Imperial College London, when they found a way to separate the cells that produce sperm from the remaining testicular tissue.
Using a sophisticated sequencing technique, the group was able to decipher the fertility code—a milestone in epigenetics, the discipline that deals with potentially inheritable modifications regulating the activity of genes.
A surprising and intriguing outcome of the study occurred when the research team found that the code does not function correctly in men who suffer from extremely low sperm production, technically termed cryptozoospermia. This revealed a previously unknown cause of male infertility and suggests new therapeutic approaches that require further research.
More information: Lara M. Siebert-Kuss et al, Genome-wide DNA methylation changes in human spermatogenesis, The American Journal of Human Genetics (2024). DOI: 10.1016/j.ajhg.2024.04.017