Early stages of breast cancer could soon be diagnosed from blood samples
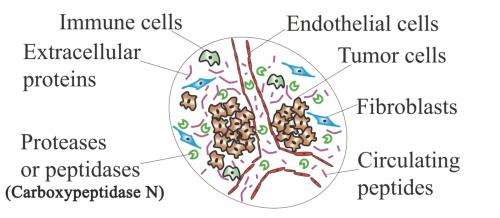
What could someday be the first blood test for the early detection of breast cancer was shown in preliminary studies to successfully identify the presence of breast cancer cells from serum biomarkers, say the Houston Methodist Research Institute scientists who are developing the technology.
With a New York University Cancer Institute colleague, the researchers report in an upcoming Clinical Chemistry (now online) that the mixture of free-floating blood proteins created by the enzyme carboxypeptidase N accurately predicted the presence of early-stage breast cancer tissue in mice and in a small population of human patients.
"In this paper we link the catalytic activity of carboxypeptidase N to tumor progression in clinical samples from breast cancer patients and a breast cancer animal model," said biomedical engineer Tony Hu, Ph.D., who led the project. "Our results indicate that circulating peptides generated by CPN can serve as clear signatures of early disease onset and progression."
The technology is not yet available to the public, and may not be for years. More extensive clinical tests are needed, and those tests are expected to begin in early 2014.
There are currently no inexpensive laboratory tests for the early detection of breast cancer, providing the impetus for researchers around the world to invent them.
"What we are trying to create is a non-invasive test that profiles what's going on at a tissue site without having to do a biopsy or costly imaging," Hu said. "We think this could be better for patients and—if we are successful—a lot cheaper than the technology that exists. While there's more to the cost of administering a test than materials alone, right now those materials only cost about $10 per test."
CPN is an enzyme that modifies proteins after the proteins are first created. Past studies have only shown the enzyme is more active in lung cancer patients. The present report in Clinical Chemistry is the first to show CPN isn't merely more active in breast cancer patients, but there's more of it.
The technology being developed by Hu's group combines nanotechnology and advanced mass spectrometry to separate and detect extremely low levels of small proteins (peptides) created by CPN. These peptides are believed to originate in or near cancerous cells, eventually making their way into the bloodstream.
In animal models and human biopsies, Hu's group first determined the presence of breast cancer tissue, characterized each sample's stage of development, and looked at how much CPN was being expressed. Blood samples were also taken from each individual.
Blood serum proteins were separated on a nanoporous silica chip dotted with four nanometer holes, which captured and isolated smaller proteins for spectrographic analysis. Using MALDI-TOF mass spectrometry, the researchers analyzed what remained for the light signatures of six peptides known to be created by CPN.
The researchers compared the stages of breast cancer tissue development in previously diagnosed patients to the presence of CPN-created peptides in their blood. They found all six peptides were at detectably higher levels, as early as breast cancer's first pathologic stage. (That stage is defined as having cancerous cells and a tumor of 2 cm or smaller, or no tumor at all.) The researchers also found that CPN peptides were at detectably higher levels in the blood of mice, compared to controls, just two weeks after the introduction of breast cancer tissue.
Interestingly, CPN activity dropped significantly over time in mice over the eight week study period, suggesting the blood test as currently configured may not work as well in detecting later stages of breast cancer. Hu said he plans to investigate the phenomenon.
"Even at the eighth week, CPN activity was still significantly higher than baseline," Hu said. "However, we suspect the activity of different enzymes goes up and down as the disease progresses. We will be looking at how we might add known and future biomarkers to the blood test to increase its robustness and accuracy."
Current means for the early detection of breast cancer are expensive and are not generally recommend for prevention by the American Cancer Society. Rather, the society recommends healthy women age 40 and older have a mammogram every year and work with their doctors to assess their individual risks of developing the disease. Prior to age 40, the society recommends women have a clinical breast exam whenever they visit their doctors, or else every three years.
More information: "Circulating Proteolytic Products of Carboxypeptidase N for Early Detection of Breast Cancer" Clinical Chemistry, DOI: 10.1373/clinchem.2013.211953