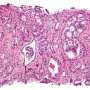
Two new genes involved in the more aggressive prostate cancer

A study by the Columbia University Nova York, in collaboration with the Catalan Institute of Oncology , Belvitge Biomedical Research Institute (ICO-IDIBELL) has identified two new genes that lead to more aggressive forms of prostate cancer. The work done by Alvaro Aytes under the direction of Cory Abate-Shen , director of the Herbert Irving Comprehensive Cancer Center of the Columbia University, has been published in the latest issue of Cancer Cell.
Prostate cancer
Prostate cancer is the most common in men in Europe( accounts for 20% of all male tumors). The incidence is about 60 new cases per 100,000 inhabitants per year.
A tumor closely associated with old age, and most cases is diagnosed between 70 and 80 years. The progressive aging of the population has made it one of the tumors that has increased in recent decades. Survival is quite high: according to Oncology Master Plan, 84 % of patients are alive five years after diagnosis.
Identify high-risk patients
In a significant proportion of patients the prostate tumor has a not very aggressive behavior, which does not compromise the health and quality of life of the affected. Also, being a tumor that usually occurs in old age, often the person dies with the tumor but not as a result of this.
It is therefore necessary to develop tools to predict which prostate tumors are clinically relevant and potentially lethal. This would open the door to customize treatments and avoid certain therapies and , therefore, avoid side effects to patients that do not need it, further reducing healthcare costs.
Two new genes in the spotlight
The study published in Cancer Cell identified two genes, the FOXM1 and CENPF, if they are abnormally activated simultaneously, lead to more aggressive and life threatening forms of prostate cancer .
A new feature of the study is that computer algorithms were used to generate networks of interactions between molecules that are generated specifically in prostate cancer .
Currently they have launched preclinical studies to determine which treatments or combinations of drugs are more effective in combating abnormal activation of FOXM1 and CENPF genes. Also, in the near future, identifying the presence or absence of these biomarkers in an individual patient will provide a more effective and with fewer side effects individualized treatment.